Study run-b2
Study informations
100 subnetworks in total page | file
190 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 3178-4-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1651 | 1.236e-02 | 2.366e-02 | 4.959e-02 |
|---|
| IPC-NIBC-129 | 0.2387 | 1.253e-01 | 8.119e-02 | 4.931e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2330 | 1.405e-02 | 8.015e-03 | 1.390e-01 |
|---|
| Loi_GPL570 | 0.1988 | 7.487e-02 | 5.091e-02 | 3.155e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1653 | 1.688e-02 | 4.170e-04 | 1.928e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.3273 | 1.041e-01 | 4.889e-02 | 4.936e-01 |
|---|
| Schmidt | 0.2201 | 1.510e-02 | 2.108e-02 | 8.481e-02 |
|---|
| Sotiriou | 0.3004 | 1.040e-01 | 7.675e-02 | 5.709e-01 |
|---|
| Zhang | 0.2493 | 6.750e-03 | 1.004e-02 | 2.901e-02 |
|---|
Expression data for subnetwork 3178-4-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Zhang |
Subnetwork structure for each dataset
- Desmedt
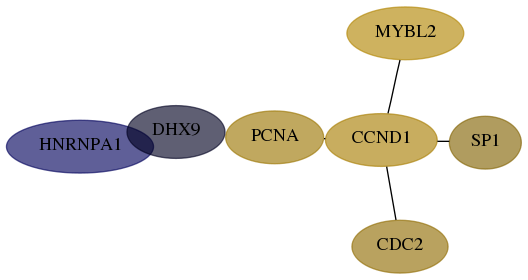
Score for each gene in subnetwork 3178-4-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Zhang |
|---|
| sp1 |   | 14 | 4 | 11 | 6 | 0.054 | 0.186 | 0.147 | 0.242 | 0.125 | 0.202 | 0.284 | 0.227 | 0.091 |
|---|
| pcna |   | 9 | 7 | 56 | 42 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.123 |
|---|
| hnrnpa1 |   | 1 | 57 | 79 | 83 | -0.021 | 0.209 | 0.088 | -0.020 | 0.087 | 0.082 | 0.036 | 0.089 | 0.083 |
|---|
| mybl2 |   | 9 | 7 | 29 | 13 | 0.162 | 0.237 | 0.166 | 0.079 | 0.124 | 0.275 | 0.202 | 0.182 | 0.074 |
|---|
| cdc2 |   | 83 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.297 |
|---|
| dhx9 |   | 1 | 57 | 79 | 83 | -0.004 | 0.165 | 0.099 | -0.094 | 0.103 | 0.127 | 0.155 | 0.135 | 0.007 |
|---|
| ccnd1 |   | 76 | 2 | 1 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.144 |
|---|
GO Enrichment output for subnetwork 3178-4-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 0.01029669 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 5.331E-05 | 0.04086901 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.03501589 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02801271 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 7.758E-05 | 0.02973854 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| regulation of chromosome organization | GO:0033044 |  | 1.517E-04 | 0.03171266 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.325E-07 | 2.034E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 1.44E-06 | 1.759E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 2.159E-06 | 1.758E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.028E-06 | 2.46E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.723E-05 | 8.42E-03 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 1.953E-05 | 7.951E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.726E-05 | 9.514E-03 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 3.313E-05 | 0.01011676 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 4.657E-05 | 0.01264064 |
|---|
| positive regulation of nuclear division | GO:0051785 |  | 4.657E-05 | 0.01137658 |
|---|
| regulation of chromosome organization | GO:0033044 |  | 5.028E-05 | 0.01116722 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.366E-06 | 3.287E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.005E-06 | 2.412E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.007E-06 | 2.411E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 5.61E-06 | 3.374E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.399E-05 | 0.01154394 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 2.718E-05 | 0.01089971 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.416E-05 | 0.01174072 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 4.611E-05 | 0.01386668 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.983E-05 | 0.01599496 |
|---|
| positive regulation of nuclear division | GO:0051785 |  | 6.48E-05 | 0.01559094 |
|---|
| fat cell differentiation | GO:0045444 |  | 6.997E-05 | 0.01530341 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.325E-07 | 2.034E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 1.44E-06 | 1.759E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 2.159E-06 | 1.758E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.028E-06 | 2.46E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.723E-05 | 8.42E-03 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 1.953E-05 | 7.951E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.726E-05 | 9.514E-03 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 3.313E-05 | 0.01011676 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 4.657E-05 | 0.01264064 |
|---|
| positive regulation of nuclear division | GO:0051785 |  | 4.657E-05 | 0.01137658 |
|---|
| regulation of chromosome organization | GO:0033044 |  | 5.028E-05 | 0.01116722 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.366E-06 | 3.287E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.005E-06 | 2.412E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.007E-06 | 2.411E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 5.61E-06 | 3.374E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.399E-05 | 0.01154394 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 2.718E-05 | 0.01089971 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.416E-05 | 0.01174072 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 4.611E-05 | 0.01386668 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.983E-05 | 0.01599496 |
|---|
| positive regulation of nuclear division | GO:0051785 |  | 6.48E-05 | 0.01559094 |
|---|
| fat cell differentiation | GO:0045444 |  | 6.997E-05 | 0.01530341 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.366E-06 | 3.287E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.005E-06 | 2.412E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.007E-06 | 2.411E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 5.61E-06 | 3.374E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.399E-05 | 0.01154394 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 2.718E-05 | 0.01089971 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.416E-05 | 0.01174072 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 4.611E-05 | 0.01386668 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.983E-05 | 0.01599496 |
|---|
| positive regulation of nuclear division | GO:0051785 |  | 6.48E-05 | 0.01559094 |
|---|
| fat cell differentiation | GO:0045444 |  | 6.997E-05 | 0.01530341 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 0.01029669 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 5.331E-05 | 0.04086901 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.03501589 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02801271 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 7.758E-05 | 0.02973854 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| regulation of chromosome organization | GO:0033044 |  | 1.517E-04 | 0.03171266 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 0.01029669 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 5.331E-05 | 0.04086901 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.03501589 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02801271 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 7.758E-05 | 0.02973854 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| regulation of chromosome organization | GO:0033044 |  | 1.517E-04 | 0.03171266 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 0.01029669 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 5.331E-05 | 0.04086901 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.03501589 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02801271 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 7.758E-05 | 0.02973854 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| regulation of chromosome organization | GO:0033044 |  | 1.517E-04 | 0.03171266 |
|---|



