Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 3978-4-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1811 | 2.097e-02 | 8.861e-03 | 1.042e-01 |
|---|
| IPC-NIBC-129 | 0.2027 | 7.221e-02 | 6.081e-02 | 3.319e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2113 | 3.042e-02 | 2.244e-02 | 3.096e-01 |
|---|
| Loi_GPL570 | 0.2439 | 6.169e-02 | 8.044e-02 | 2.651e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1577 | 6.622e-03 | 2.822e-03 | 8.132e-02 |
|---|
| Pawitan_GPL96-GPL97 | 0.3468 | 1.069e-01 | 5.932e-02 | 5.166e-01 |
|---|
| Schmidt | 0.2469 | 4.088e-02 | 4.297e-02 | 2.417e-01 |
|---|
| Sotiriou | 0.2390 | 1.252e-01 | 5.769e-02 | 6.416e-01 |
|---|
| Wang | 0.1791 | 6.839e-02 | 5.220e-02 | 2.283e-01 |
|---|
| Zhang | 0.2487 | 4.876e-03 | 2.283e-03 | 2.585e-02 |
|---|
Expression data for subnetwork 3978-4-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
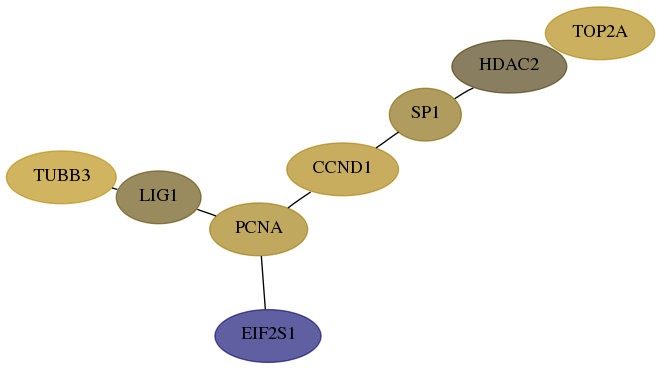
Score for each gene in subnetwork 3978-4-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| pcna |   | 39 | 3 | 8 | 3 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.162 | 0.123 |
|---|
| tubb3 |   | 1 | 90 | 147 | 153 | 0.183 | 0.221 | 0.105 | 0.017 | 0.138 | 0.334 | 0.055 | 0.119 | -0.023 | -0.083 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| sp1 |   | 31 | 5 | 16 | 8 | 0.054 | 0.186 | 0.147 | 0.242 | 0.125 | 0.202 | 0.284 | 0.227 | 0.194 | 0.091 |
|---|
| lig1 |   | 1 | 90 | 147 | 153 | 0.023 | 0.117 | 0.134 | 0.142 | 0.116 | 0.090 | 0.125 | 0.212 | 0.070 | 0.122 |
|---|
| hdac2 |   | 16 | 10 | 64 | 39 | 0.012 | 0.245 | 0.030 | 0.077 | 0.070 | 0.183 | 0.222 | -0.058 | 0.086 | 0.133 |
|---|
| top2a |   | 22 | 7 | 70 | 46 | 0.147 | 0.183 | 0.185 | 0.100 | 0.206 | 0.309 | 0.261 | 0.183 | 0.072 | 0.234 |
|---|
| eif2s1 |   | 13 | 12 | 8 | 4 | -0.031 | 0.151 | 0.253 | 0.182 | 0.098 | 0.270 | 0.233 | 0.038 | 0.122 | 0.239 |
|---|
GO Enrichment output for subnetwork 3978-4-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.275E-07 | 5.234E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 5.688E-06 | 6.541E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 5.947E-06 | 4.56E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 5.947E-06 | 3.42E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 8.829E-06 | 4.061E-03 |
|---|
| DNA topological change | GO:0006265 |  | 8.917E-06 | 3.418E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.166E-05 | 3.832E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.248E-05 | 3.587E-03 |
|---|
| V(D)J recombination | GO:0033151 |  | 2.137E-05 | 5.461E-03 |
|---|
| DNA ligation | GO:0006266 |  | 2.137E-05 | 4.915E-03 |
|---|
| natural killer cell mediated immunity | GO:0002228 |  | 2.137E-05 | 4.468E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.394E-08 | 8.291E-05 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.04E-06 | 1.27E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 1.661E-06 | 1.353E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 1.661E-06 | 1.014E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 1.755E-06 | 8.574E-04 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 2.491E-06 | 1.014E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 2.491E-06 | 8.693E-04 |
|---|
| chromosome segregation | GO:0007059 |  | 2.866E-06 | 8.751E-04 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 4.031E-06 | 1.094E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.648E-06 | 1.135E-03 |
|---|
| DNA topological change | GO:0006265 |  | 4.648E-06 | 1.032E-03 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.575E-08 | 1.341E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.706E-06 | 2.052E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 2.313E-06 | 1.855E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 2.313E-06 | 1.391E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.313E-06 | 1.113E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 2.727E-06 | 1.094E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.469E-06 | 1.192E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 4.697E-06 | 1.413E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.472E-06 | 1.73E-03 |
|---|
| DNA topological change | GO:0006265 |  | 6.472E-06 | 1.557E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 6.605E-06 | 1.445E-03 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.394E-08 | 8.291E-05 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.04E-06 | 1.27E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 1.661E-06 | 1.353E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 1.661E-06 | 1.014E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 1.755E-06 | 8.574E-04 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 2.491E-06 | 1.014E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 2.491E-06 | 8.693E-04 |
|---|
| chromosome segregation | GO:0007059 |  | 2.866E-06 | 8.751E-04 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 4.031E-06 | 1.094E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.648E-06 | 1.135E-03 |
|---|
| DNA topological change | GO:0006265 |  | 4.648E-06 | 1.032E-03 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.575E-08 | 1.341E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.706E-06 | 2.052E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 2.313E-06 | 1.855E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 2.313E-06 | 1.391E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.313E-06 | 1.113E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 2.727E-06 | 1.094E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.469E-06 | 1.192E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 4.697E-06 | 1.413E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.472E-06 | 1.73E-03 |
|---|
| DNA topological change | GO:0006265 |  | 6.472E-06 | 1.557E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 6.605E-06 | 1.445E-03 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.575E-08 | 1.341E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.706E-06 | 2.052E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 2.313E-06 | 1.855E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 2.313E-06 | 1.391E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.313E-06 | 1.113E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 2.727E-06 | 1.094E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.469E-06 | 1.192E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 4.697E-06 | 1.413E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.472E-06 | 1.73E-03 |
|---|
| DNA topological change | GO:0006265 |  | 6.472E-06 | 1.557E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 6.605E-06 | 1.445E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.275E-07 | 5.234E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 5.688E-06 | 6.541E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 5.947E-06 | 4.56E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 5.947E-06 | 3.42E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 8.829E-06 | 4.061E-03 |
|---|
| DNA topological change | GO:0006265 |  | 8.917E-06 | 3.418E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.166E-05 | 3.832E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.248E-05 | 3.587E-03 |
|---|
| V(D)J recombination | GO:0033151 |  | 2.137E-05 | 5.461E-03 |
|---|
| DNA ligation | GO:0006266 |  | 2.137E-05 | 4.915E-03 |
|---|
| natural killer cell mediated immunity | GO:0002228 |  | 2.137E-05 | 4.468E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.275E-07 | 5.234E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 5.688E-06 | 6.541E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 5.947E-06 | 4.56E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 5.947E-06 | 3.42E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 8.829E-06 | 4.061E-03 |
|---|
| DNA topological change | GO:0006265 |  | 8.917E-06 | 3.418E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.166E-05 | 3.832E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.248E-05 | 3.587E-03 |
|---|
| V(D)J recombination | GO:0033151 |  | 2.137E-05 | 5.461E-03 |
|---|
| DNA ligation | GO:0006266 |  | 2.137E-05 | 4.915E-03 |
|---|
| natural killer cell mediated immunity | GO:0002228 |  | 2.137E-05 | 4.468E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.275E-07 | 5.234E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 5.688E-06 | 6.541E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 5.947E-06 | 4.56E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 5.947E-06 | 3.42E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 8.829E-06 | 4.061E-03 |
|---|
| DNA topological change | GO:0006265 |  | 8.917E-06 | 3.418E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.166E-05 | 3.832E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.248E-05 | 3.587E-03 |
|---|
| V(D)J recombination | GO:0033151 |  | 2.137E-05 | 5.461E-03 |
|---|
| DNA ligation | GO:0006266 |  | 2.137E-05 | 4.915E-03 |
|---|
| natural killer cell mediated immunity | GO:0002228 |  | 2.137E-05 | 4.468E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.275E-07 | 5.234E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 5.688E-06 | 6.541E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 5.947E-06 | 4.56E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 5.947E-06 | 3.42E-03 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 8.829E-06 | 4.061E-03 |
|---|
| DNA topological change | GO:0006265 |  | 8.917E-06 | 3.418E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.166E-05 | 3.832E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.248E-05 | 3.587E-03 |
|---|
| V(D)J recombination | GO:0033151 |  | 2.137E-05 | 5.461E-03 |
|---|
| DNA ligation | GO:0006266 |  | 2.137E-05 | 4.915E-03 |
|---|
| natural killer cell mediated immunity | GO:0002228 |  | 2.137E-05 | 4.468E-03 |
|---|



