Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 22907-2-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1116 | 1.167e-02 | 3.992e-03 | 6.584e-02 |
|---|
| IPC-NIBC-129 | 0.1909 | 4.011e-02 | 3.112e-02 | 1.945e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2394 | 3.834e-02 | 2.939e-02 | 3.654e-01 |
|---|
| Loi_GPL570 | 0.2902 | 4.528e-02 | 6.052e-02 | 2.077e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1603 | 8.999e-02 | 8.623e-02 | 7.774e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.3194 | 5.077e-02 | 2.282e-02 | 3.123e-01 |
|---|
| Schmidt | 0.2699 | 2.636e-02 | 2.799e-02 | 1.696e-01 |
|---|
| Sotiriou | 0.2669 | 1.202e-01 | 5.441e-02 | 6.281e-01 |
|---|
| Wang | 0.2397 | 2.401e-02 | 1.363e-02 | 1.015e-01 |
|---|
| Zhang | 0.3167 | 7.763e-03 | 3.836e-03 | 3.798e-02 |
|---|
Expression data for subnetwork 22907-2-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
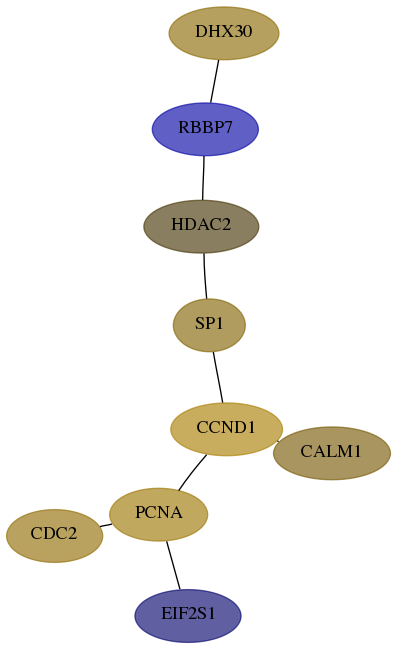
Score for each gene in subnetwork 22907-2-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| pcna |   | 39 | 3 | 8 | 3 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.162 | 0.123 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| sp1 |   | 31 | 5 | 16 | 8 | 0.054 | 0.186 | 0.147 | 0.242 | 0.125 | 0.202 | 0.284 | 0.227 | 0.194 | 0.091 |
|---|
| rbbp7 |   | 1 | 90 | 64 | 74 | -0.113 | 0.089 | 0.051 | 0.076 | 0.106 | -0.020 | 0.126 | 0.138 | 0.115 | 0.168 |
|---|
| calm1 |   | 16 | 10 | 35 | 19 | 0.038 | 0.023 | 0.171 | 0.277 | 0.113 | 0.007 | 0.171 | 0.112 | 0.087 | 0.162 |
|---|
| dhx30 |   | 1 | 90 | 64 | 74 | 0.066 | -0.039 | 0.122 | -0.072 | 0.099 | 0.326 | 0.065 | 0.160 | -0.028 | 0.015 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| hdac2 |   | 16 | 10 | 64 | 39 | 0.012 | 0.245 | 0.030 | 0.077 | 0.070 | 0.183 | 0.222 | -0.058 | 0.086 | 0.133 |
|---|
| eif2s1 |   | 13 | 12 | 8 | 4 | -0.031 | 0.151 | 0.253 | 0.182 | 0.098 | 0.270 | 0.233 | 0.038 | 0.122 | 0.239 |
|---|
GO Enrichment output for subnetwork 22907-2-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.052E-05 | 0.04719789 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.876E-05 | 0.03307007 |
|---|
| histone deacetylation | GO:0016575 |  | 7.51E-05 | 0.05757426 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 1.43E-04 | 0.0821973 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.633E-04 | 0.07509739 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.633E-04 | 0.06258116 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.633E-04 | 0.053641 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 2.078E-04 | 0.0597566 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.847E-04 | 0.07274741 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 3.425E-04 | 0.07876481 |
|---|
| fat cell differentiation | GO:0045444 |  | 3.733E-04 | 0.07805731 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 4.743E-06 | 0.01158756 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 5.206E-06 | 6.359E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 7.112E-06 | 5.791E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.326E-05 | 8.101E-03 |
|---|
| histone deacetylation | GO:0016575 |  | 4.959E-05 | 0.02422913 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.665E-05 | 0.02306542 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 6.417E-05 | 0.02239674 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 8.954E-05 | 0.02734297 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 8.954E-05 | 0.02430486 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 1.088E-04 | 0.02657171 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.528E-04 | 0.03392728 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 6.605E-06 | 0.01589185 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.526E-06 | 0.01025628 |
|---|
| establishment of spindle localization | GO:0051293 |  | 9.903E-06 | 7.942E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.847E-05 | 0.01110731 |
|---|
| histone deacetylation | GO:0016575 |  | 6.9E-05 | 0.03320339 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 7.882E-05 | 0.03160619 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 8.928E-05 | 0.03068753 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.121E-04 | 0.03372757 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.245E-04 | 0.03329423 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 1.513E-04 | 0.03639384 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.961E-04 | 0.04290239 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 4.743E-06 | 0.01158756 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 5.206E-06 | 6.359E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 7.112E-06 | 5.791E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.326E-05 | 8.101E-03 |
|---|
| histone deacetylation | GO:0016575 |  | 4.959E-05 | 0.02422913 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.665E-05 | 0.02306542 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 6.417E-05 | 0.02239674 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 8.954E-05 | 0.02734297 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 8.954E-05 | 0.02430486 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 1.088E-04 | 0.02657171 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.528E-04 | 0.03392728 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 6.605E-06 | 0.01589185 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.526E-06 | 0.01025628 |
|---|
| establishment of spindle localization | GO:0051293 |  | 9.903E-06 | 7.942E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.847E-05 | 0.01110731 |
|---|
| histone deacetylation | GO:0016575 |  | 6.9E-05 | 0.03320339 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 7.882E-05 | 0.03160619 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 8.928E-05 | 0.03068753 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.121E-04 | 0.03372757 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.245E-04 | 0.03329423 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 1.513E-04 | 0.03639384 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.961E-04 | 0.04290239 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 6.605E-06 | 0.01589185 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.526E-06 | 0.01025628 |
|---|
| establishment of spindle localization | GO:0051293 |  | 9.903E-06 | 7.942E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.847E-05 | 0.01110731 |
|---|
| histone deacetylation | GO:0016575 |  | 6.9E-05 | 0.03320339 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 7.882E-05 | 0.03160619 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 8.928E-05 | 0.03068753 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.121E-04 | 0.03372757 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.245E-04 | 0.03329423 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 1.513E-04 | 0.03639384 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.961E-04 | 0.04290239 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.052E-05 | 0.04719789 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.876E-05 | 0.03307007 |
|---|
| histone deacetylation | GO:0016575 |  | 7.51E-05 | 0.05757426 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 1.43E-04 | 0.0821973 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.633E-04 | 0.07509739 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.633E-04 | 0.06258116 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.633E-04 | 0.053641 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 2.078E-04 | 0.0597566 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.847E-04 | 0.07274741 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 3.425E-04 | 0.07876481 |
|---|
| fat cell differentiation | GO:0045444 |  | 3.733E-04 | 0.07805731 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.052E-05 | 0.04719789 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.876E-05 | 0.03307007 |
|---|
| histone deacetylation | GO:0016575 |  | 7.51E-05 | 0.05757426 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 1.43E-04 | 0.0821973 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.633E-04 | 0.07509739 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.633E-04 | 0.06258116 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.633E-04 | 0.053641 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 2.078E-04 | 0.0597566 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.847E-04 | 0.07274741 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 3.425E-04 | 0.07876481 |
|---|
| fat cell differentiation | GO:0045444 |  | 3.733E-04 | 0.07805731 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.052E-05 | 0.04719789 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.876E-05 | 0.03307007 |
|---|
| histone deacetylation | GO:0016575 |  | 7.51E-05 | 0.05757426 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 1.43E-04 | 0.0821973 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.633E-04 | 0.07509739 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.633E-04 | 0.06258116 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.633E-04 | 0.053641 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 2.078E-04 | 0.0597566 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.847E-04 | 0.07274741 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 3.425E-04 | 0.07876481 |
|---|
| fat cell differentiation | GO:0045444 |  | 3.733E-04 | 0.07805731 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.052E-05 | 0.04719789 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.876E-05 | 0.03307007 |
|---|
| histone deacetylation | GO:0016575 |  | 7.51E-05 | 0.05757426 |
|---|
| negative regulation of cell cycle process | GO:0010948 |  | 1.43E-04 | 0.0821973 |
|---|
| protein amino acid deacetylation | GO:0006476 |  | 1.633E-04 | 0.07509739 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.633E-04 | 0.06258116 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.633E-04 | 0.053641 |
|---|
| regulation of mitotic metaphase/anaphase transition | GO:0030071 |  | 2.078E-04 | 0.0597566 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.847E-04 | 0.07274741 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 3.425E-04 | 0.07876481 |
|---|
| fat cell differentiation | GO:0045444 |  | 3.733E-04 | 0.07805731 |
|---|



