Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 1164-1-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1472 | 1.372e-02 | 4.980e-03 | 7.484e-02 |
|---|
| IPC-NIBC-129 | 0.1700 | 9.892e-02 | 8.685e-02 | 4.285e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2317 | 5.760e-02 | 4.714e-02 | 4.772e-01 |
|---|
| Loi_GPL570 | 0.2188 | 1.281e-01 | 1.573e-01 | 4.499e-01 |
|---|
| Loi_GPL96-GPL97 | 0.2128 | 2.473e-02 | 1.637e-02 | 3.345e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.2473 | 5.033e-02 | 2.257e-02 | 3.103e-01 |
|---|
| Schmidt | 0.1915 | 2.113e-02 | 2.254e-02 | 1.407e-01 |
|---|
| Sotiriou | 0.2809 | 5.160e-02 | 1.626e-02 | 3.712e-01 |
|---|
| Wang | 0.1891 | 5.766e-02 | 4.203e-02 | 2.008e-01 |
|---|
| Zhang | 0.3175 | 2.605e-02 | 1.487e-02 | 1.009e-01 |
|---|
Expression data for subnetwork 1164-1-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
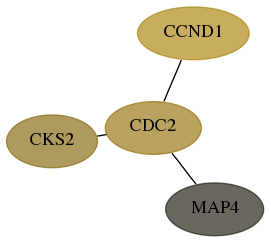
Score for each gene in subnetwork 1164-1-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| cks2 |   | 4 | 27 | 151 | 122 | 0.050 | 0.233 | 0.170 | 0.103 | 0.186 | 0.167 | 0.263 | 0.151 | 0.145 | 0.284 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| map4 |   | 1 | 90 | 191 | 192 | 0.002 | 0.046 | 0.067 | -0.043 | 0.034 | -0.058 | -0.020 | 0.043 | 0.108 | -0.074 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
GO Enrichment output for subnetwork 1164-1-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 0.01161803 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 0.01041131 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 3.913E-05 | 0.02999691 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 4.513E-05 | 0.02594897 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.156E-05 | 0.02371572 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 7.338E-05 | 0.02813021 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.005E-05 | 0.02958817 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.084E-04 | 0.03116708 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.182E-04 | 0.0302111 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 1.391E-04 | 0.03199278 |
|---|
| spindle organization | GO:0007051 |  | 1.859E-04 | 0.03886893 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.755E-06 | 4.287E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.648E-06 | 5.677E-03 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 2.252E-05 | 0.0183422 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 2.533E-05 | 0.01547246 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.144E-05 | 0.01536384 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 4.563E-05 | 0.0185802 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.371E-05 | 0.01874416 |
|---|
| fat cell differentiation | GO:0045444 |  | 6.244E-05 | 0.01906649 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.704E-05 | 0.01819896 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 7.675E-05 | 0.01875013 |
|---|
| spindle organization | GO:0007051 |  | 8.185E-05 | 0.01817751 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.878E-06 | 6.924E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.472E-06 | 7.785E-03 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 2.767E-05 | 0.022193 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 3.135E-05 | 0.01885862 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.94E-05 | 0.01895864 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 5.318E-05 | 0.02132391 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 6.9E-05 | 0.02371671 |
|---|
| fat cell differentiation | GO:0045444 |  | 8.068E-05 | 0.02426601 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 8.687E-05 | 0.02322232 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 9.327E-05 | 0.02244178 |
|---|
| spindle organization | GO:0007051 |  | 1.139E-04 | 0.02490264 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.755E-06 | 4.287E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.648E-06 | 5.677E-03 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 2.252E-05 | 0.0183422 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 2.533E-05 | 0.01547246 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.144E-05 | 0.01536384 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 4.563E-05 | 0.0185802 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.371E-05 | 0.01874416 |
|---|
| fat cell differentiation | GO:0045444 |  | 6.244E-05 | 0.01906649 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.704E-05 | 0.01819896 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 7.675E-05 | 0.01875013 |
|---|
| spindle organization | GO:0007051 |  | 8.185E-05 | 0.01817751 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.878E-06 | 6.924E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.472E-06 | 7.785E-03 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 2.767E-05 | 0.022193 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 3.135E-05 | 0.01885862 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.94E-05 | 0.01895864 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 5.318E-05 | 0.02132391 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 6.9E-05 | 0.02371671 |
|---|
| fat cell differentiation | GO:0045444 |  | 8.068E-05 | 0.02426601 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 8.687E-05 | 0.02322232 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 9.327E-05 | 0.02244178 |
|---|
| spindle organization | GO:0007051 |  | 1.139E-04 | 0.02490264 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.878E-06 | 6.924E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.472E-06 | 7.785E-03 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 2.767E-05 | 0.022193 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 3.135E-05 | 0.01885862 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 3.94E-05 | 0.01895864 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 5.318E-05 | 0.02132391 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 6.9E-05 | 0.02371671 |
|---|
| fat cell differentiation | GO:0045444 |  | 8.068E-05 | 0.02426601 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 8.687E-05 | 0.02322232 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 9.327E-05 | 0.02244178 |
|---|
| spindle organization | GO:0007051 |  | 1.139E-04 | 0.02490264 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 0.01161803 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 0.01041131 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 3.913E-05 | 0.02999691 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 4.513E-05 | 0.02594897 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.156E-05 | 0.02371572 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 7.338E-05 | 0.02813021 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.005E-05 | 0.02958817 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.084E-04 | 0.03116708 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.182E-04 | 0.0302111 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 1.391E-04 | 0.03199278 |
|---|
| spindle organization | GO:0007051 |  | 1.859E-04 | 0.03886893 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 0.01161803 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 0.01041131 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 3.913E-05 | 0.02999691 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 4.513E-05 | 0.02594897 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.156E-05 | 0.02371572 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 7.338E-05 | 0.02813021 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.005E-05 | 0.02958817 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.084E-04 | 0.03116708 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.182E-04 | 0.0302111 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 1.391E-04 | 0.03199278 |
|---|
| spindle organization | GO:0007051 |  | 1.859E-04 | 0.03886893 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 0.01161803 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 0.01041131 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 3.913E-05 | 0.02999691 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 4.513E-05 | 0.02594897 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.156E-05 | 0.02371572 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 7.338E-05 | 0.02813021 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.005E-05 | 0.02958817 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.084E-04 | 0.03116708 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.182E-04 | 0.0302111 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 1.391E-04 | 0.03199278 |
|---|
| spindle organization | GO:0007051 |  | 1.859E-04 | 0.03886893 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 0.01161803 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 0.01041131 |
|---|
| negative regulation of microtubule depolymerization | GO:0007026 |  | 3.913E-05 | 0.02999691 |
|---|
| negative regulation of microtubule polymerization or depolymerization | GO:0031111 |  | 4.513E-05 | 0.02594897 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.156E-05 | 0.02371572 |
|---|
| regulation of microtubule polymerization or depolymerization | GO:0031110 |  | 7.338E-05 | 0.02813021 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.005E-05 | 0.02958817 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.084E-04 | 0.03116708 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.182E-04 | 0.0302111 |
|---|
| regulation of microtubule cytoskeleton organization | GO:0070507 |  | 1.391E-04 | 0.03199278 |
|---|
| spindle organization | GO:0007051 |  | 1.859E-04 | 0.03886893 |
|---|



