Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 11339-0-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1456 | 1.374e-02 | 4.987e-03 | 7.490e-02 |
|---|
| IPC-NIBC-129 | 0.0995 | 4.337e-02 | 3.404e-02 | 2.097e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2316 | 2.192e-01 | 2.165e-01 | 8.730e-01 |
|---|
| Loi_GPL570 | 0.2841 | 1.039e-01 | 1.298e-01 | 3.895e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1696 | 2.627e-02 | 1.773e-02 | 3.526e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.3557 | 8.066e-02 | 4.132e-02 | 4.320e-01 |
|---|
| Schmidt | 0.2076 | 2.845e-02 | 3.015e-02 | 1.806e-01 |
|---|
| Sotiriou | 0.2621 | 1.038e-01 | 4.414e-02 | 5.801e-01 |
|---|
| Wang | 0.1762 | 7.193e-02 | 5.565e-02 | 2.370e-01 |
|---|
| Zhang | 0.2746 | 4.195e-03 | 1.930e-03 | 2.281e-02 |
|---|
Expression data for subnetwork 11339-0-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
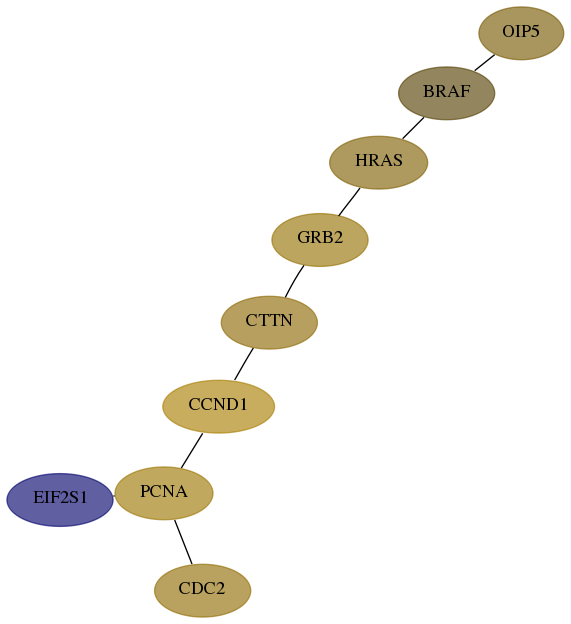
Score for each gene in subnetwork 11339-0-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| pcna |   | 39 | 3 | 8 | 3 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.162 | 0.123 |
|---|
| cttn |   | 5 | 20 | 8 | 9 | 0.067 | 0.065 | 0.123 | 0.238 | 0.126 | 0.216 | 0.069 | 0.115 | 0.129 | 0.025 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| hras |   | 5 | 20 | 51 | 36 | 0.049 | -0.030 | 0.094 | 0.210 | 0.160 | 0.245 | 0.086 | 0.208 | 0.001 | 0.003 |
|---|
| oip5 |   | 1 | 90 | 192 | 193 | 0.040 | 0.082 | 0.085 | 0.226 | 0.100 | 0.140 | 0.164 | 0.059 | 0.120 | 0.065 |
|---|
| braf |   | 1 | 90 | 192 | 193 | 0.017 | -0.109 | 0.073 | 0.006 | 0.072 | 0.157 | 0.092 | 0.071 | 0.070 | 0.033 |
|---|
| eif2s1 |   | 13 | 12 | 8 | 4 | -0.031 | 0.151 | 0.253 | 0.182 | 0.098 | 0.270 | 0.233 | 0.038 | 0.122 | 0.239 |
|---|
| grb2 |   | 6 | 17 | 8 | 7 | 0.084 | -0.013 | 0.086 | -0.031 | 0.162 | 0.065 | 0.158 | 0.127 | -0.097 | 0.194 |
|---|
GO Enrichment output for subnetwork 11339-0-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 0.03063632 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 0.02143316 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 0.0190408 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 2.484E-05 | 0.0142806 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 3.987E-05 | 0.01833977 |
|---|
| response to light stimulus | GO:0009416 |  | 1.032E-04 | 0.03955791 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.06E-04 | 0.03481319 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.06E-04 | 0.03046155 |
|---|
| Ras protein signal transduction | GO:0007265 |  | 1.165E-04 | 0.02977394 |
|---|
| visual learning | GO:0008542 |  | 1.349E-04 | 0.03103525 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 1.507E-04 | 0.03151511 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 7.565E-06 | 0.01848231 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 7.565E-06 | 9.241E-03 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 7.565E-06 | 6.161E-03 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 1.215E-05 | 7.421E-03 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.454E-05 | 0.01199054 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.234E-05 | 0.01316802 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 4.604E-05 | 0.01606841 |
|---|
| visual learning | GO:0008542 |  | 4.604E-05 | 0.01405986 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.114E-05 | 0.01388186 |
|---|
| cell aging | GO:0007569 |  | 6.214E-05 | 0.01517999 |
|---|
| visual behavior | GO:0007632 |  | 6.803E-05 | 0.01510946 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.053E-05 | 0.02534487 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.053E-05 | 0.01267243 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 1.053E-05 | 8.448E-03 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 1.692E-05 | 0.01017554 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.067E-05 | 9.946E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 4.501E-05 | 0.01804905 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 6.407E-05 | 0.02202076 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.407E-05 | 0.01926816 |
|---|
| visual learning | GO:0008542 |  | 6.407E-05 | 0.01712726 |
|---|
| cell aging | GO:0007569 |  | 7.862E-05 | 0.01891589 |
|---|
| visual behavior | GO:0007632 |  | 9.465E-05 | 0.02070182 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 7.565E-06 | 0.01848231 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 7.565E-06 | 9.241E-03 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 7.565E-06 | 6.161E-03 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 1.215E-05 | 7.421E-03 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.454E-05 | 0.01199054 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.234E-05 | 0.01316802 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 4.604E-05 | 0.01606841 |
|---|
| visual learning | GO:0008542 |  | 4.604E-05 | 0.01405986 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.114E-05 | 0.01388186 |
|---|
| cell aging | GO:0007569 |  | 6.214E-05 | 0.01517999 |
|---|
| visual behavior | GO:0007632 |  | 6.803E-05 | 0.01510946 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.053E-05 | 0.02534487 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.053E-05 | 0.01267243 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 1.053E-05 | 8.448E-03 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 1.692E-05 | 0.01017554 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.067E-05 | 9.946E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 4.501E-05 | 0.01804905 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 6.407E-05 | 0.02202076 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.407E-05 | 0.01926816 |
|---|
| visual learning | GO:0008542 |  | 6.407E-05 | 0.01712726 |
|---|
| cell aging | GO:0007569 |  | 7.862E-05 | 0.01891589 |
|---|
| visual behavior | GO:0007632 |  | 9.465E-05 | 0.02070182 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.053E-05 | 0.02534487 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.053E-05 | 0.01267243 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 1.053E-05 | 8.448E-03 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 1.692E-05 | 0.01017554 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.067E-05 | 9.946E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 4.501E-05 | 0.01804905 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 6.407E-05 | 0.02202076 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.407E-05 | 0.01926816 |
|---|
| visual learning | GO:0008542 |  | 6.407E-05 | 0.01712726 |
|---|
| cell aging | GO:0007569 |  | 7.862E-05 | 0.01891589 |
|---|
| visual behavior | GO:0007632 |  | 9.465E-05 | 0.02070182 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 0.03063632 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 0.02143316 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 0.0190408 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 2.484E-05 | 0.0142806 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 3.987E-05 | 0.01833977 |
|---|
| response to light stimulus | GO:0009416 |  | 1.032E-04 | 0.03955791 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.06E-04 | 0.03481319 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.06E-04 | 0.03046155 |
|---|
| Ras protein signal transduction | GO:0007265 |  | 1.165E-04 | 0.02977394 |
|---|
| visual learning | GO:0008542 |  | 1.349E-04 | 0.03103525 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 1.507E-04 | 0.03151511 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 0.03063632 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 0.02143316 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 0.0190408 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 2.484E-05 | 0.0142806 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 3.987E-05 | 0.01833977 |
|---|
| response to light stimulus | GO:0009416 |  | 1.032E-04 | 0.03955791 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.06E-04 | 0.03481319 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.06E-04 | 0.03046155 |
|---|
| Ras protein signal transduction | GO:0007265 |  | 1.165E-04 | 0.02977394 |
|---|
| visual learning | GO:0008542 |  | 1.349E-04 | 0.03103525 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 1.507E-04 | 0.03151511 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 0.03063632 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 0.02143316 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 0.0190408 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 2.484E-05 | 0.0142806 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 3.987E-05 | 0.01833977 |
|---|
| response to light stimulus | GO:0009416 |  | 1.032E-04 | 0.03955791 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.06E-04 | 0.03481319 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.06E-04 | 0.03046155 |
|---|
| Ras protein signal transduction | GO:0007265 |  | 1.165E-04 | 0.02977394 |
|---|
| visual learning | GO:0008542 |  | 1.349E-04 | 0.03103525 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 1.507E-04 | 0.03151511 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 0.03063632 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 0.02143316 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 0.0190408 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 2.484E-05 | 0.0142806 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 3.987E-05 | 0.01833977 |
|---|
| response to light stimulus | GO:0009416 |  | 1.032E-04 | 0.03955791 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.06E-04 | 0.03481319 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.06E-04 | 0.03046155 |
|---|
| Ras protein signal transduction | GO:0007265 |  | 1.165E-04 | 0.02977394 |
|---|
| visual learning | GO:0008542 |  | 1.349E-04 | 0.03103525 |
|---|
| regulation of neuronal synaptic plasticity | GO:0048168 |  | 1.507E-04 | 0.03151511 |
|---|



