Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 10915-0-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1783 | 1.237e-02 | 4.322e-03 | 6.896e-02 |
|---|
| IPC-NIBC-129 | 0.2294 | 1.276e-01 | 1.157e-01 | 5.169e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2366 | 1.793e-02 | 1.208e-02 | 2.051e-01 |
|---|
| Loi_GPL570 | 0.1982 | 1.661e-01 | 1.994e-01 | 5.320e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1676 | 7.388e-03 | 3.271e-03 | 9.304e-02 |
|---|
| Pawitan_GPL96-GPL97 | 0.2812 | 6.471e-02 | 3.115e-02 | 3.718e-01 |
|---|
| Schmidt | 0.1714 | 1.367e-02 | 1.473e-02 | 9.620e-02 |
|---|
| Sotiriou | 0.3082 | 1.072e-01 | 4.619e-02 | 5.905e-01 |
|---|
| Wang | 0.1613 | 9.258e-02 | 7.643e-02 | 2.852e-01 |
|---|
| Zhang | 0.2947 | 1.477e-02 | 7.872e-03 | 6.412e-02 |
|---|
Expression data for subnetwork 10915-0-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
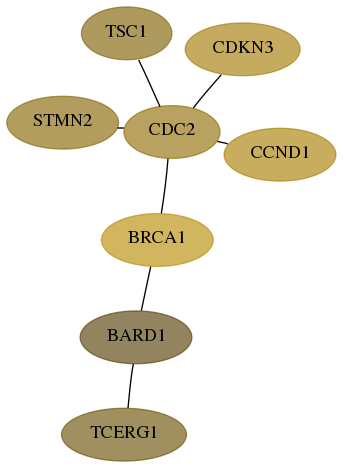
Score for each gene in subnetwork 10915-0-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| cdkn3 |   | 18 | 9 | 26 | 16 | 0.114 | 0.269 | 0.199 | 0.136 | 0.193 | 0.323 | 0.186 | 0.198 | 0.134 | 0.271 |
|---|
| tcerg1 |   | 1 | 90 | 165 | 174 | 0.029 | 0.245 | 0.043 | 0.052 | 0.094 | 0.015 | 0.141 | 0.159 | 0.142 | -0.016 |
|---|
| stmn2 |   | 19 | 8 | 32 | 18 | 0.057 | 0.106 | 0.005 | 0.052 | 0.110 | 0.006 | 0.034 | 0.103 | 0.075 | 0.113 |
|---|
| brca1 |   | 5 | 20 | 104 | 78 | 0.192 | 0.052 | 0.112 | 0.022 | 0.125 | 0.113 | 0.077 | 0.168 | 0.155 | 0.064 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| tsc1 |   | 31 | 5 | 26 | 15 | 0.049 | 0.012 | -0.055 | 0.072 | 0.055 | -0.196 | 0.006 | 0.010 | 0.043 | -0.007 |
|---|
| bard1 |   | 1 | 90 | 165 | 174 | 0.016 | 0.161 | 0.164 | 0.042 | 0.137 | 0.153 | 0.022 | 0.142 | 0.103 | 0.122 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
GO Enrichment output for subnetwork 10915-0-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.119E-07 | 2.574E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.003E-06 | 3.453E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 4.314E-06 | 3.308E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 4.314E-06 | 2.481E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 4.314E-06 | 1.985E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 1.936E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 6.469E-06 | 2.126E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 6.469E-06 | 1.86E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 6.469E-06 | 1.653E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 6.469E-06 | 1.488E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 1.893E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 5.069E-08 | 1.238E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.362E-06 | 1.664E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.151E-06 | 1.752E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.151E-06 | 1.314E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.617E-06 | 1.278E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.226E-06 | 1.313E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.226E-06 | 1.126E-03 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 3.226E-06 | 9.851E-04 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 4.515E-06 | 1.226E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 4.515E-06 | 1.103E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.018E-06 | 1.337E-03 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 6.86E-08 | 1.65E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.841E-06 | 2.215E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.644E-06 | 2.12E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.644E-06 | 1.59E-03 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.644E-06 | 1.272E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.537E-06 | 1.418E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.964E-06 | 1.363E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.964E-06 | 1.192E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 5.548E-06 | 1.483E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 5.548E-06 | 1.335E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 7.395E-06 | 1.618E-03 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 5.069E-08 | 1.238E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.362E-06 | 1.664E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.151E-06 | 1.752E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.151E-06 | 1.314E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.617E-06 | 1.278E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.226E-06 | 1.313E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.226E-06 | 1.126E-03 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 3.226E-06 | 9.851E-04 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 4.515E-06 | 1.226E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 4.515E-06 | 1.103E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.018E-06 | 1.337E-03 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 6.86E-08 | 1.65E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.841E-06 | 2.215E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.644E-06 | 2.12E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.644E-06 | 1.59E-03 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.644E-06 | 1.272E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.537E-06 | 1.418E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.964E-06 | 1.363E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.964E-06 | 1.192E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 5.548E-06 | 1.483E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 5.548E-06 | 1.335E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 7.395E-06 | 1.618E-03 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 6.86E-08 | 1.65E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.841E-06 | 2.215E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.644E-06 | 2.12E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.644E-06 | 1.59E-03 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.644E-06 | 1.272E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.537E-06 | 1.418E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.964E-06 | 1.363E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.964E-06 | 1.192E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 5.548E-06 | 1.483E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 5.548E-06 | 1.335E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 7.395E-06 | 1.618E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.119E-07 | 2.574E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.003E-06 | 3.453E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 4.314E-06 | 3.308E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 4.314E-06 | 2.481E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 4.314E-06 | 1.985E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 1.936E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 6.469E-06 | 2.126E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 6.469E-06 | 1.86E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 6.469E-06 | 1.653E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 6.469E-06 | 1.488E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 1.893E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.119E-07 | 2.574E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.003E-06 | 3.453E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 4.314E-06 | 3.308E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 4.314E-06 | 2.481E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 4.314E-06 | 1.985E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 1.936E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 6.469E-06 | 2.126E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 6.469E-06 | 1.86E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 6.469E-06 | 1.653E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 6.469E-06 | 1.488E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 1.893E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.119E-07 | 2.574E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.003E-06 | 3.453E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 4.314E-06 | 3.308E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 4.314E-06 | 2.481E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 4.314E-06 | 1.985E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 1.936E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 6.469E-06 | 2.126E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 6.469E-06 | 1.86E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 6.469E-06 | 1.653E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 6.469E-06 | 1.488E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 1.893E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.119E-07 | 2.574E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.003E-06 | 3.453E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 4.314E-06 | 3.308E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 4.314E-06 | 2.481E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 4.314E-06 | 1.985E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.051E-06 | 1.936E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 6.469E-06 | 2.126E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 6.469E-06 | 1.86E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 6.469E-06 | 1.653E-03 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 6.469E-06 | 1.488E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.053E-06 | 1.893E-03 |
|---|



