Study run-b2
Study informations
100 subnetworks in total page | file
190 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 4172-5-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1533 | 1.409e-02 | 2.658e-02 | 5.593e-02 |
|---|
| IPC-NIBC-129 | 0.1930 | 1.189e-01 | 7.620e-02 | 4.721e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2268 | 3.533e-02 | 2.422e-02 | 3.178e-01 |
|---|
| Loi_GPL570 | 0.2031 | 6.436e-02 | 4.151e-02 | 2.805e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1888 | 2.552e-02 | 1.340e-03 | 2.988e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.3578 | 1.607e-01 | 8.869e-02 | 6.400e-01 |
|---|
| Schmidt | 0.2309 | 3.536e-02 | 4.784e-02 | 1.931e-01 |
|---|
| Sotiriou | 0.2466 | 7.027e-02 | 5.082e-02 | 4.401e-01 |
|---|
| Zhang | 0.2087 | 4.060e-03 | 6.027e-03 | 1.868e-02 |
|---|
Expression data for subnetwork 4172-5-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Zhang |
Subnetwork structure for each dataset
- Desmedt
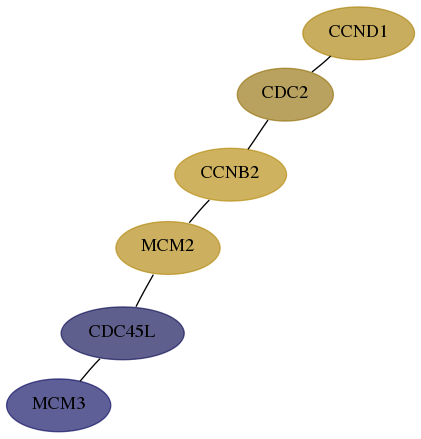
Score for each gene in subnetwork 4172-5-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Zhang |
|---|
| mcm3 |   | 1 | 57 | 140 | 141 | -0.020 | 0.173 | 0.113 | 0.084 | 0.110 | 0.255 | 0.171 | 0.156 | -0.003 |
|---|
| ccnb2 |   | 5 | 13 | 58 | 45 | 0.164 | 0.176 | 0.204 | 0.123 | 0.172 | 0.366 | 0.240 | 0.169 | 0.211 |
|---|
| cdc2 |   | 83 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.297 |
|---|
| mcm2 |   | 2 | 35 | 90 | 82 | 0.151 | 0.254 | 0.188 | 0.128 | 0.154 | 0.267 | 0.224 | 0.200 | 0.012 |
|---|
| cdc45l |   | 2 | 35 | 113 | 108 | -0.014 | 0.244 | 0.143 | 0.181 | 0.118 | 0.227 | 0.262 | 0.050 | 0.189 |
|---|
| ccnd1 |   | 76 | 2 | 1 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.144 |
|---|
GO Enrichment output for subnetwork 4172-5-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 5.878E-11 | 1.352E-07 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 3.961E-09 | 4.555E-06 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.941E-06 | 3.788E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 6.586E-06 | 3.787E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 8.466E-06 | 3.894E-03 |
|---|
| thymus development | GO:0048538 |  | 8.466E-06 | 3.245E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 1.293E-05 | 4.247E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 1.832E-05 | 5.268E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 2.137E-05 | 5.461E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.817E-05 | 6.478E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 3.191E-05 | 6.673E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 5.739E-12 | 1.402E-08 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 8.495E-10 | 1.038E-06 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.993E-06 | 1.623E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 1.993E-06 | 1.217E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 3.202E-06 | 1.564E-03 |
|---|
| thymus development | GO:0048538 |  | 3.913E-06 | 1.593E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 4.695E-06 | 1.638E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 7.466E-06 | 2.28E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 9.667E-06 | 2.624E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 1.35E-05 | 3.298E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.35E-05 | 2.998E-03 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 6.68E-12 | 1.607E-08 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 7.743E-10 | 9.315E-07 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.221E-06 | 1.781E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 2.221E-06 | 1.336E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 3.568E-06 | 1.717E-03 |
|---|
| thymus development | GO:0048538 |  | 4.36E-06 | 1.748E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 5.231E-06 | 1.798E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 8.318E-06 | 2.502E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 1.077E-05 | 2.879E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.354E-05 | 3.257E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 1.504E-05 | 3.29E-03 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 5.739E-12 | 1.402E-08 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 8.495E-10 | 1.038E-06 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.993E-06 | 1.623E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 1.993E-06 | 1.217E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 3.202E-06 | 1.564E-03 |
|---|
| thymus development | GO:0048538 |  | 3.913E-06 | 1.593E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 4.695E-06 | 1.638E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 7.466E-06 | 2.28E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 9.667E-06 | 2.624E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 1.35E-05 | 3.298E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.35E-05 | 2.998E-03 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 6.68E-12 | 1.607E-08 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 7.743E-10 | 9.315E-07 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.221E-06 | 1.781E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 2.221E-06 | 1.336E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 3.568E-06 | 1.717E-03 |
|---|
| thymus development | GO:0048538 |  | 4.36E-06 | 1.748E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 5.231E-06 | 1.798E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 8.318E-06 | 2.502E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 1.077E-05 | 2.879E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.354E-05 | 3.257E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 1.504E-05 | 3.29E-03 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 6.68E-12 | 1.607E-08 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 7.743E-10 | 9.315E-07 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.221E-06 | 1.781E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 2.221E-06 | 1.336E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 3.568E-06 | 1.717E-03 |
|---|
| thymus development | GO:0048538 |  | 4.36E-06 | 1.748E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 5.231E-06 | 1.798E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 8.318E-06 | 2.502E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 1.077E-05 | 2.879E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.354E-05 | 3.257E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 1.504E-05 | 3.29E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 5.878E-11 | 1.352E-07 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 3.961E-09 | 4.555E-06 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.941E-06 | 3.788E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 6.586E-06 | 3.787E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 8.466E-06 | 3.894E-03 |
|---|
| thymus development | GO:0048538 |  | 8.466E-06 | 3.245E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 1.293E-05 | 4.247E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 1.832E-05 | 5.268E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 2.137E-05 | 5.461E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.817E-05 | 6.478E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 3.191E-05 | 6.673E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 5.878E-11 | 1.352E-07 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 3.961E-09 | 4.555E-06 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.941E-06 | 3.788E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 6.586E-06 | 3.787E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 8.466E-06 | 3.894E-03 |
|---|
| thymus development | GO:0048538 |  | 8.466E-06 | 3.245E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 1.293E-05 | 4.247E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 1.832E-05 | 5.268E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 2.137E-05 | 5.461E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.817E-05 | 6.478E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 3.191E-05 | 6.673E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| DNA replication initiation | GO:0006270 |  | 5.878E-11 | 1.352E-07 |
|---|
| DNA-dependent DNA replication | GO:0006261 |  | 3.961E-09 | 4.555E-06 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.941E-06 | 3.788E-03 |
|---|
| negative regulation of DNA replication initiation | GO:0032297 |  | 6.586E-06 | 3.787E-03 |
|---|
| regulation of DNA replication initiation | GO:0030174 |  | 8.466E-06 | 3.894E-03 |
|---|
| thymus development | GO:0048538 |  | 8.466E-06 | 3.245E-03 |
|---|
| T cell homeostasis | GO:0043029 |  | 1.293E-05 | 4.247E-03 |
|---|
| lymphocyte homeostasis | GO:0002260 |  | 1.832E-05 | 5.268E-03 |
|---|
| DNA unwinding during replication | GO:0006268 |  | 2.137E-05 | 5.461E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.817E-05 | 6.478E-03 |
|---|
| DNA duplex unwinding | GO:0032508 |  | 3.191E-05 | 6.673E-03 |
|---|



