Study run-b2
Study informations
100 subnetworks in total page | file
190 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 152-1-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1836 | 1.605e-02 | 2.987e-02 | 6.303e-02 |
|---|
| IPC-NIBC-129 | 0.1556 | 4.952e-02 | 2.666e-02 | 2.000e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2205 | 7.396e-02 | 5.826e-02 | 5.399e-01 |
|---|
| Loi_GPL570 | 0.2736 | 6.824e-02 | 4.493e-02 | 2.937e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1713 | 8.699e-03 | 6.000e-05 | 8.529e-02 |
|---|
| Pawitan_GPL96-GPL97 | 0.3289 | 9.217e-02 | 4.136e-02 | 4.553e-01 |
|---|
| Schmidt | 0.2267 | 1.580e-02 | 2.201e-02 | 8.876e-02 |
|---|
| Sotiriou | 0.2975 | 9.411e-02 | 6.906e-02 | 5.363e-01 |
|---|
| Zhang | 0.2605 | 6.573e-03 | 9.771e-03 | 2.835e-02 |
|---|
Expression data for subnetwork 152-1-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Zhang |
Subnetwork structure for each dataset
- Desmedt
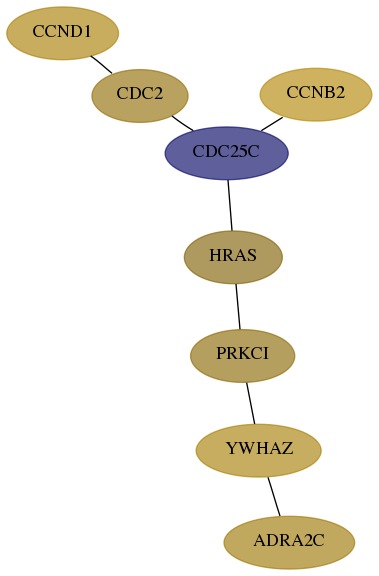
Score for each gene in subnetwork 152-1-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Zhang |
|---|
| ccnb2 |   | 5 | 13 | 58 | 45 | 0.164 | 0.176 | 0.204 | 0.123 | 0.172 | 0.366 | 0.240 | 0.169 | 0.211 |
|---|
| cdc2 |   | 83 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.297 |
|---|
| hras |   | 3 | 25 | 58 | 50 | 0.049 | -0.030 | 0.094 | 0.210 | 0.160 | 0.245 | 0.086 | 0.208 | 0.003 |
|---|
| cdc25c |   | 1 | 57 | 61 | 65 | -0.025 | 0.187 | 0.084 | 0.267 | 0.070 | 0.125 | 0.204 | 0.250 | 0.054 |
|---|
| ywhaz |   | 9 | 7 | 17 | 9 | 0.124 | 0.089 | 0.087 | 0.200 | 0.141 | 0.199 | 0.056 | 0.123 | 0.103 |
|---|
| prkci |   | 2 | 35 | 61 | 53 | 0.065 | 0.104 | 0.012 | 0.170 | 0.112 | -0.170 | 0.196 | 0.128 | 0.124 |
|---|
| ccnd1 |   | 76 | 2 | 1 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.144 |
|---|
| adra2c |   | 2 | 35 | 61 | 53 | 0.100 | 0.086 | 0.058 | 0.087 | 0.055 | 0.090 | 0.095 | -0.051 | 0.120 |
|---|
GO Enrichment output for subnetwork 152-1-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 6.283E-09 | 1.445E-05 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 1.033E-07 | 1.188E-04 |
|---|
| fructose metabolic process | GO:0006000 |  | 3.497E-07 | 2.681E-04 |
|---|
| alditol metabolic process | GO:0019400 |  | 7.103E-07 | 4.084E-04 |
|---|
| sperm motility | GO:0030317 |  | 1.258E-06 | 5.788E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 2.265E-06 | 8.683E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 3.07E-06 | 1.009E-03 |
|---|
| polyol metabolic process | GO:0019751 |  | 3.699E-06 | 1.064E-03 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 5.203E-06 | 1.33E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 7.841E-06 | 1.803E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.175E-05 | 2.458E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 9.081E-10 | 2.219E-06 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 2.591E-08 | 3.165E-05 |
|---|
| fructose metabolic process | GO:0006000 |  | 5.069E-08 | 4.128E-05 |
|---|
| alditol metabolic process | GO:0019400 |  | 2.346E-07 | 1.433E-04 |
|---|
| sperm motility | GO:0030317 |  | 2.955E-07 | 1.444E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 4.468E-07 | 1.819E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 6.425E-07 | 2.242E-04 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 1.031E-06 | 3.149E-04 |
|---|
| polyol metabolic process | GO:0019751 |  | 1.273E-06 | 3.457E-04 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.617E-06 | 6.392E-04 |
|---|
| photoreceptor cell development | GO:0042461 |  | 3.226E-06 | 7.164E-04 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 1.229E-09 | 2.958E-06 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 3.507E-08 | 4.219E-05 |
|---|
| fructose metabolic process | GO:0006000 |  | 6.86E-08 | 5.502E-05 |
|---|
| alditol metabolic process | GO:0019400 |  | 2.473E-07 | 1.487E-04 |
|---|
| sperm motility | GO:0030317 |  | 2.809E-07 | 1.352E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 6.044E-07 | 2.424E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 8.69E-07 | 2.987E-04 |
|---|
| polyol metabolic process | GO:0019751 |  | 1.295E-06 | 3.896E-04 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 1.394E-06 | 3.728E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.644E-06 | 6.361E-04 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.537E-06 | 7.736E-04 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 9.081E-10 | 2.219E-06 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 2.591E-08 | 3.165E-05 |
|---|
| fructose metabolic process | GO:0006000 |  | 5.069E-08 | 4.128E-05 |
|---|
| alditol metabolic process | GO:0019400 |  | 2.346E-07 | 1.433E-04 |
|---|
| sperm motility | GO:0030317 |  | 2.955E-07 | 1.444E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 4.468E-07 | 1.819E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 6.425E-07 | 2.242E-04 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 1.031E-06 | 3.149E-04 |
|---|
| polyol metabolic process | GO:0019751 |  | 1.273E-06 | 3.457E-04 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.617E-06 | 6.392E-04 |
|---|
| photoreceptor cell development | GO:0042461 |  | 3.226E-06 | 7.164E-04 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 1.229E-09 | 2.958E-06 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 3.507E-08 | 4.219E-05 |
|---|
| fructose metabolic process | GO:0006000 |  | 6.86E-08 | 5.502E-05 |
|---|
| alditol metabolic process | GO:0019400 |  | 2.473E-07 | 1.487E-04 |
|---|
| sperm motility | GO:0030317 |  | 2.809E-07 | 1.352E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 6.044E-07 | 2.424E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 8.69E-07 | 2.987E-04 |
|---|
| polyol metabolic process | GO:0019751 |  | 1.295E-06 | 3.896E-04 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 1.394E-06 | 3.728E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.644E-06 | 6.361E-04 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.537E-06 | 7.736E-04 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 1.229E-09 | 2.958E-06 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 3.507E-08 | 4.219E-05 |
|---|
| fructose metabolic process | GO:0006000 |  | 6.86E-08 | 5.502E-05 |
|---|
| alditol metabolic process | GO:0019400 |  | 2.473E-07 | 1.487E-04 |
|---|
| sperm motility | GO:0030317 |  | 2.809E-07 | 1.352E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 6.044E-07 | 2.424E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 8.69E-07 | 2.987E-04 |
|---|
| polyol metabolic process | GO:0019751 |  | 1.295E-06 | 3.896E-04 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 1.394E-06 | 3.728E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.644E-06 | 6.361E-04 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.537E-06 | 7.736E-04 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 6.283E-09 | 1.445E-05 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 1.033E-07 | 1.188E-04 |
|---|
| fructose metabolic process | GO:0006000 |  | 3.497E-07 | 2.681E-04 |
|---|
| alditol metabolic process | GO:0019400 |  | 7.103E-07 | 4.084E-04 |
|---|
| sperm motility | GO:0030317 |  | 1.258E-06 | 5.788E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 2.265E-06 | 8.683E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 3.07E-06 | 1.009E-03 |
|---|
| polyol metabolic process | GO:0019751 |  | 3.699E-06 | 1.064E-03 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 5.203E-06 | 1.33E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 7.841E-06 | 1.803E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.175E-05 | 2.458E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 6.283E-09 | 1.445E-05 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 1.033E-07 | 1.188E-04 |
|---|
| fructose metabolic process | GO:0006000 |  | 3.497E-07 | 2.681E-04 |
|---|
| alditol metabolic process | GO:0019400 |  | 7.103E-07 | 4.084E-04 |
|---|
| sperm motility | GO:0030317 |  | 1.258E-06 | 5.788E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 2.265E-06 | 8.683E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 3.07E-06 | 1.009E-03 |
|---|
| polyol metabolic process | GO:0019751 |  | 3.699E-06 | 1.064E-03 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 5.203E-06 | 1.33E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 7.841E-06 | 1.803E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.175E-05 | 2.458E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| polyol catabolic process | GO:0046174 |  | 6.283E-09 | 1.445E-05 |
|---|
| cellular carbohydrate catabolic process | GO:0044275 |  | 1.033E-07 | 1.188E-04 |
|---|
| fructose metabolic process | GO:0006000 |  | 3.497E-07 | 2.681E-04 |
|---|
| alditol metabolic process | GO:0019400 |  | 7.103E-07 | 4.084E-04 |
|---|
| sperm motility | GO:0030317 |  | 1.258E-06 | 5.788E-04 |
|---|
| hexose biosynthetic process | GO:0019319 |  | 2.265E-06 | 8.683E-04 |
|---|
| monosaccharide biosynthetic process | GO:0046364 |  | 3.07E-06 | 1.009E-03 |
|---|
| polyol metabolic process | GO:0019751 |  | 3.699E-06 | 1.064E-03 |
|---|
| alcohol biosynthetic process | GO:0046165 |  | 5.203E-06 | 1.33E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 7.841E-06 | 1.803E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.175E-05 | 2.458E-03 |
|---|



