Study run-b2
Study informations
100 subnetworks in total page | file
190 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 10615-0-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1802 | 1.231e-02 | 2.358e-02 | 4.942e-02 |
|---|
| IPC-NIBC-129 | 0.1498 | 1.406e-01 | 9.323e-02 | 5.402e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2332 | 8.285e-02 | 6.661e-02 | 5.782e-01 |
|---|
| Loi_GPL570 | 0.1893 | 1.625e-01 | 1.418e-01 | 5.440e-01 |
|---|
| Loi_GPL96-GPL97 | 0.2517 | 9.851e-03 | 8.700e-05 | 1.004e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.2639 | 6.091e-02 | 2.339e-02 | 3.380e-01 |
|---|
| Schmidt | 0.1635 | 1.948e-02 | 2.695e-02 | 1.095e-01 |
|---|
| Sotiriou | 0.2843 | 2.397e-02 | 1.660e-02 | 1.799e-01 |
|---|
| Zhang | 0.2984 | 1.933e-02 | 2.858e-02 | 7.081e-02 |
|---|
Expression data for subnetwork 10615-0-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Zhang |
Subnetwork structure for each dataset
- Desmedt
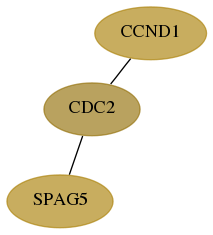
Score for each gene in subnetwork 10615-0-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Zhang |
|---|
| cdc2 |   | 83 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.297 |
|---|
| spag5 |   | 1 | 57 | 138 | 139 | 0.127 | 0.245 | 0.169 | 0.063 | 0.186 | 0.087 | 0.212 | 0.133 | 0.144 |
|---|
| ccnd1 |   | 76 | 2 | 1 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.144 |
|---|
GO Enrichment output for subnetwork 10615-0-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.06E-06 | 4.738E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.175E-05 | 0.01351809 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.056E-05 | 0.01575907 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 2.476E-05 | 0.0142351 |
|---|
| fat cell differentiation | GO:0045444 |  | 2.7E-05 | 0.01242146 |
|---|
| spindle organization | GO:0007051 |  | 4.252E-05 | 0.01629948 |
|---|
| response to calcium ion | GO:0051592 |  | 7.613E-05 | 0.0250132 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 8.809E-05 | 0.02532629 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 9.655E-05 | 0.02467367 |
|---|
| response to UV | GO:0009411 |  | 1.009E-04 | 0.02321214 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.242E-04 | 0.0259753 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.646E-07 | 1.624E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 4.506E-06 | 5.504E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 7.704E-06 | 6.273E-03 |
|---|
| fat cell differentiation | GO:0045444 |  | 8.959E-06 | 5.472E-03 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 9.622E-06 | 4.701E-03 |
|---|
| spindle organization | GO:0007051 |  | 1.175E-05 | 4.785E-03 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 2.45E-05 | 8.549E-03 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.669E-05 | 8.152E-03 |
|---|
| response to calcium ion | GO:0051592 |  | 2.783E-05 | 7.554E-03 |
|---|
| response to UV | GO:0009411 |  | 3.512E-05 | 8.581E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.774E-05 | 8.382E-03 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.256E-07 | 2.227E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.647E-06 | 6.794E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.903E-06 | 7.942E-03 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.158E-05 | 6.968E-03 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.247E-05 | 6.002E-03 |
|---|
| spindle organization | GO:0007051 |  | 1.636E-05 | 6.561E-03 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.41E-05 | 0.01172081 |
|---|
| response to calcium ion | GO:0051592 |  | 3.561E-05 | 0.01071057 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 3.716E-05 | 9.934E-03 |
|---|
| response to UV | GO:0009411 |  | 4.537E-05 | 0.01091679 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.253E-05 | 0.01149032 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.646E-07 | 1.624E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 4.506E-06 | 5.504E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 7.704E-06 | 6.273E-03 |
|---|
| fat cell differentiation | GO:0045444 |  | 8.959E-06 | 5.472E-03 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 9.622E-06 | 4.701E-03 |
|---|
| spindle organization | GO:0007051 |  | 1.175E-05 | 4.785E-03 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 2.45E-05 | 8.549E-03 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.669E-05 | 8.152E-03 |
|---|
| response to calcium ion | GO:0051592 |  | 2.783E-05 | 7.554E-03 |
|---|
| response to UV | GO:0009411 |  | 3.512E-05 | 8.581E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 3.774E-05 | 8.382E-03 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.256E-07 | 2.227E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.647E-06 | 6.794E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.903E-06 | 7.942E-03 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.158E-05 | 6.968E-03 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.247E-05 | 6.002E-03 |
|---|
| spindle organization | GO:0007051 |  | 1.636E-05 | 6.561E-03 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.41E-05 | 0.01172081 |
|---|
| response to calcium ion | GO:0051592 |  | 3.561E-05 | 0.01071057 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 3.716E-05 | 9.934E-03 |
|---|
| response to UV | GO:0009411 |  | 4.537E-05 | 0.01091679 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.253E-05 | 0.01149032 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.256E-07 | 2.227E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 5.647E-06 | 6.794E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 9.903E-06 | 7.942E-03 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.158E-05 | 6.968E-03 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.247E-05 | 6.002E-03 |
|---|
| spindle organization | GO:0007051 |  | 1.636E-05 | 6.561E-03 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.41E-05 | 0.01172081 |
|---|
| response to calcium ion | GO:0051592 |  | 3.561E-05 | 0.01071057 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 3.716E-05 | 9.934E-03 |
|---|
| response to UV | GO:0009411 |  | 4.537E-05 | 0.01091679 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 5.253E-05 | 0.01149032 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.06E-06 | 4.738E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.175E-05 | 0.01351809 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.056E-05 | 0.01575907 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 2.476E-05 | 0.0142351 |
|---|
| fat cell differentiation | GO:0045444 |  | 2.7E-05 | 0.01242146 |
|---|
| spindle organization | GO:0007051 |  | 4.252E-05 | 0.01629948 |
|---|
| response to calcium ion | GO:0051592 |  | 7.613E-05 | 0.0250132 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 8.809E-05 | 0.02532629 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 9.655E-05 | 0.02467367 |
|---|
| response to UV | GO:0009411 |  | 1.009E-04 | 0.02321214 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.242E-04 | 0.0259753 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.06E-06 | 4.738E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.175E-05 | 0.01351809 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.056E-05 | 0.01575907 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 2.476E-05 | 0.0142351 |
|---|
| fat cell differentiation | GO:0045444 |  | 2.7E-05 | 0.01242146 |
|---|
| spindle organization | GO:0007051 |  | 4.252E-05 | 0.01629948 |
|---|
| response to calcium ion | GO:0051592 |  | 7.613E-05 | 0.0250132 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 8.809E-05 | 0.02532629 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 9.655E-05 | 0.02467367 |
|---|
| response to UV | GO:0009411 |  | 1.009E-04 | 0.02321214 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.242E-04 | 0.0259753 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 2.06E-06 | 4.738E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 1.175E-05 | 0.01351809 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 2.056E-05 | 0.01575907 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 2.476E-05 | 0.0142351 |
|---|
| fat cell differentiation | GO:0045444 |  | 2.7E-05 | 0.01242146 |
|---|
| spindle organization | GO:0007051 |  | 4.252E-05 | 0.01629948 |
|---|
| response to calcium ion | GO:0051592 |  | 7.613E-05 | 0.0250132 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 8.809E-05 | 0.02532629 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 9.655E-05 | 0.02467367 |
|---|
| response to UV | GO:0009411 |  | 1.009E-04 | 0.02321214 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.242E-04 | 0.0259753 |
|---|



