Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 8812-10-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.2126 | 1.378e-02 | 5.006e-03 | 7.507e-02 |
|---|
| IPC-NIBC-129 | 0.2282 | 1.265e-01 | 1.146e-01 | 5.138e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2315 | 1.837e-02 | 1.244e-02 | 2.093e-01 |
|---|
| Loi_GPL570 | 0.1989 | 1.258e-01 | 1.547e-01 | 4.444e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1496 | 1.848e-03 | 4.960e-04 | 1.408e-02 |
|---|
| Pawitan_GPL96-GPL97 | 0.3199 | 5.720e-02 | 2.659e-02 | 3.406e-01 |
|---|
| Schmidt | 0.1929 | 9.881e-03 | 1.073e-02 | 7.172e-02 |
|---|
| Sotiriou | 0.3285 | 1.425e-01 | 6.930e-02 | 6.839e-01 |
|---|
| Wang | 0.1518 | 1.089e-01 | 9.356e-02 | 3.203e-01 |
|---|
| Zhang | 0.3060 | 7.699e-03 | 3.801e-03 | 3.772e-02 |
|---|
Expression data for subnetwork 8812-10-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
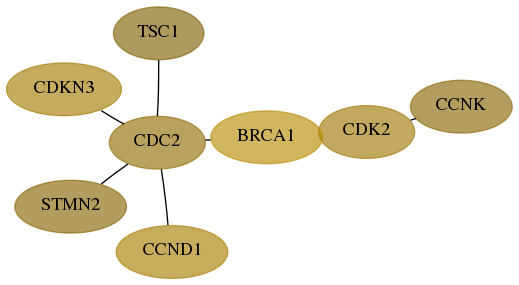
Score for each gene in subnetwork 8812-10-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| cdkn3 |   | 18 | 9 | 26 | 16 | 0.114 | 0.269 | 0.199 | 0.136 | 0.193 | 0.323 | 0.186 | 0.198 | 0.134 | 0.271 |
|---|
| stmn2 |   | 19 | 8 | 32 | 18 | 0.057 | 0.106 | 0.005 | 0.052 | 0.110 | 0.006 | 0.034 | 0.103 | 0.075 | 0.113 |
|---|
| brca1 |   | 5 | 20 | 104 | 78 | 0.192 | 0.052 | 0.112 | 0.022 | 0.125 | 0.113 | 0.077 | 0.168 | 0.155 | 0.064 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| tsc1 |   | 31 | 5 | 26 | 15 | 0.049 | 0.012 | -0.055 | 0.072 | 0.055 | -0.196 | 0.006 | 0.010 | 0.043 | -0.007 |
|---|
| ccnk |   | 1 | 90 | 104 | 112 | 0.059 | 0.082 | 0.068 | -0.043 | 0.028 | 0.157 | 0.222 | 0.093 | -0.007 | -0.087 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| cdk2 |   | 3 | 39 | 104 | 93 | 0.102 | 0.241 | 0.092 | 0.008 | 0.100 | 0.236 | 0.154 | 0.122 | 0.186 | 0.046 |
|---|
GO Enrichment output for subnetwork 8812-10-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.505E-08 | 1.036E-04 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.454E-07 | 1.672E-04 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.773E-07 | 2.893E-04 |
|---|
| interphase | GO:0051325 |  | 4.14E-07 | 2.38E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 1.793E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 5.098E-06 | 1.954E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 5.098E-06 | 1.675E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 5.098E-06 | 1.466E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 7.644E-06 | 1.758E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 7.644E-06 | 1.598E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.066E-08 | 2.604E-05 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 4.176E-08 | 5.1E-05 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 9.639E-08 | 7.849E-05 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.123E-06 | 6.856E-04 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 1.898E-06 | 9.275E-04 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 1.898E-06 | 7.729E-04 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 2.847E-06 | 9.935E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.847E-06 | 8.693E-04 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 2.847E-06 | 7.727E-04 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.847E-06 | 6.954E-04 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 3.984E-06 | 8.849E-04 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.549E-08 | 3.727E-05 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 5.575E-08 | 6.707E-05 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 1.235E-07 | 9.902E-05 |
|---|
| interphase | GO:0051325 |  | 1.4E-07 | 8.419E-05 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.498E-06 | 7.208E-04 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.313E-06 | 9.276E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.313E-06 | 7.951E-04 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.313E-06 | 6.957E-04 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.313E-06 | 6.184E-04 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.469E-06 | 8.346E-04 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.469E-06 | 7.588E-04 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.066E-08 | 2.604E-05 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 4.176E-08 | 5.1E-05 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 9.639E-08 | 7.849E-05 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.123E-06 | 6.856E-04 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 1.898E-06 | 9.275E-04 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 1.898E-06 | 7.729E-04 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 2.847E-06 | 9.935E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.847E-06 | 8.693E-04 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 2.847E-06 | 7.727E-04 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.847E-06 | 6.954E-04 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 3.984E-06 | 8.849E-04 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.549E-08 | 3.727E-05 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 5.575E-08 | 6.707E-05 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 1.235E-07 | 9.902E-05 |
|---|
| interphase | GO:0051325 |  | 1.4E-07 | 8.419E-05 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.498E-06 | 7.208E-04 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.313E-06 | 9.276E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.313E-06 | 7.951E-04 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.313E-06 | 6.957E-04 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.313E-06 | 6.184E-04 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.469E-06 | 8.346E-04 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.469E-06 | 7.588E-04 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.549E-08 | 3.727E-05 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 5.575E-08 | 6.707E-05 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 1.235E-07 | 9.902E-05 |
|---|
| interphase | GO:0051325 |  | 1.4E-07 | 8.419E-05 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 1.498E-06 | 7.208E-04 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 2.313E-06 | 9.276E-04 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 2.313E-06 | 7.951E-04 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 2.313E-06 | 6.957E-04 |
|---|
| activation of Ras GTPase activity | GO:0032856 |  | 2.313E-06 | 6.184E-04 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 3.469E-06 | 8.346E-04 |
|---|
| DNA damage response. signal transduction resulting in transcription | GO:0042772 |  | 3.469E-06 | 7.588E-04 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.505E-08 | 1.036E-04 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.454E-07 | 1.672E-04 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.773E-07 | 2.893E-04 |
|---|
| interphase | GO:0051325 |  | 4.14E-07 | 2.38E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 1.793E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 5.098E-06 | 1.954E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 5.098E-06 | 1.675E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 5.098E-06 | 1.466E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 7.644E-06 | 1.758E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 7.644E-06 | 1.598E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.505E-08 | 1.036E-04 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.454E-07 | 1.672E-04 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.773E-07 | 2.893E-04 |
|---|
| interphase | GO:0051325 |  | 4.14E-07 | 2.38E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 1.793E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 5.098E-06 | 1.954E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 5.098E-06 | 1.675E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 5.098E-06 | 1.466E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 7.644E-06 | 1.758E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 7.644E-06 | 1.598E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.505E-08 | 1.036E-04 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.454E-07 | 1.672E-04 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.773E-07 | 2.893E-04 |
|---|
| interphase | GO:0051325 |  | 4.14E-07 | 2.38E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 1.793E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 5.098E-06 | 1.954E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 5.098E-06 | 1.675E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 5.098E-06 | 1.466E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 7.644E-06 | 1.758E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 7.644E-06 | 1.598E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.505E-08 | 1.036E-04 |
|---|
| regulation of protein ubiquitination | GO:0031396 |  | 1.454E-07 | 1.672E-04 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.773E-07 | 2.893E-04 |
|---|
| interphase | GO:0051325 |  | 4.14E-07 | 2.38E-04 |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 1.793E-03 |
|---|
| negative regulation of protein ubiquitination | GO:0031397 |  | 5.098E-06 | 1.954E-03 |
|---|
| traversing start control point of mitotic cell cycle | GO:0007089 |  | 5.098E-06 | 1.675E-03 |
|---|
| regulation of centrosome cycle | GO:0046605 |  | 5.098E-06 | 1.466E-03 |
|---|
| positive regulation of Rho GTPase activity | GO:0032321 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of protein ubiquitination | GO:0031398 |  | 7.644E-06 | 1.758E-03 |
|---|
| positive regulation of DNA repair | GO:0045739 |  | 7.644E-06 | 1.598E-03 |
|---|



