Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 6502-8-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1329 | 1.280e-02 | 4.527e-03 | 7.083e-02 |
|---|
| IPC-NIBC-129 | 0.1938 | 7.536e-02 | 6.383e-02 | 3.441e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2350 | 3.624e-02 | 2.752e-02 | 3.512e-01 |
|---|
| Loi_GPL570 | 0.2405 | 8.552e-02 | 1.086e-01 | 3.389e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1702 | 4.215e-02 | 3.283e-02 | 5.103e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.3502 | 6.437e-02 | 3.094e-02 | 3.705e-01 |
|---|
| Schmidt | 0.2223 | 3.447e-02 | 3.637e-02 | 2.111e-01 |
|---|
| Sotiriou | 0.2499 | 1.028e-01 | 4.353e-02 | 5.769e-01 |
|---|
| Wang | 0.1901 | 5.666e-02 | 4.111e-02 | 1.982e-01 |
|---|
| Zhang | 0.2952 | 4.605e-03 | 2.142e-03 | 2.465e-02 |
|---|
Expression data for subnetwork 6502-8-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
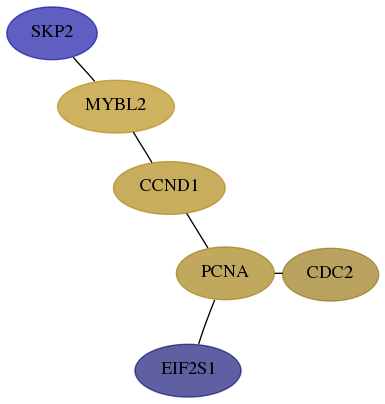
Score for each gene in subnetwork 6502-8-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| pcna |   | 39 | 3 | 8 | 3 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.162 | 0.123 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| mybl2 |   | 5 | 20 | 92 | 63 | 0.162 | 0.237 | 0.166 | 0.079 | 0.124 | 0.275 | 0.202 | 0.182 | 0.120 | 0.074 |
|---|
| skp2 |   | 1 | 90 | 128 | 134 | -0.101 | 0.121 | 0.060 | 0.145 | 0.073 | 0.188 | 0.224 | 0.031 | 0.002 | 0.000 |
|---|
| eif2s1 |   | 13 | 12 | 8 | 4 | -0.031 | 0.151 | 0.253 | 0.182 | 0.098 | 0.270 | 0.233 | 0.038 | 0.122 | 0.239 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
GO Enrichment output for subnetwork 6502-8-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 8.963E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.172E-05 | 0.02432144 |
|---|
| interphase | GO:0051325 |  | 3.399E-05 | 0.01954254 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.02801271 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02334393 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| response to calcium ion | GO:0051592 |  | 3.919E-04 | 0.08193571 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 5.749E-07 | 1.404E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 3.453E-06 | 4.218E-03 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 5.697E-06 | 4.639E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.478E-05 | 9.024E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.337E-05 | 0.01142085 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 3.993E-05 | 0.01625957 |
|---|
| fat cell differentiation | GO:0045444 |  | 4.643E-05 | 0.01620286 |
|---|
| base-excision repair | GO:0006284 |  | 4.643E-05 | 0.0141775 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 4.986E-05 | 0.01353293 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.38E-04 | 0.03370659 |
|---|
| response to calcium ion | GO:0051592 |  | 1.438E-04 | 0.0319397 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 9.435E-07 | 2.27E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.809E-06 | 5.785E-03 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 8.499E-06 | 6.816E-03 |
|---|
| interphase | GO:0051325 |  | 9.332E-06 | 5.613E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.057E-05 | 9.898E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.929E-05 | 0.01174514 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.131E-05 | 0.01763607 |
|---|
| fat cell differentiation | GO:0045444 |  | 6E-05 | 0.01804614 |
|---|
| base-excision repair | GO:0006284 |  | 6E-05 | 0.01604101 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.46E-05 | 0.01554366 |
|---|
| response to calcium ion | GO:0051592 |  | 1.839E-04 | 0.040224 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 5.749E-07 | 1.404E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 3.453E-06 | 4.218E-03 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 5.697E-06 | 4.639E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 1.478E-05 | 9.024E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.337E-05 | 0.01142085 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 3.993E-05 | 0.01625957 |
|---|
| fat cell differentiation | GO:0045444 |  | 4.643E-05 | 0.01620286 |
|---|
| base-excision repair | GO:0006284 |  | 4.643E-05 | 0.0141775 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 4.986E-05 | 0.01353293 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.38E-04 | 0.03370659 |
|---|
| response to calcium ion | GO:0051592 |  | 1.438E-04 | 0.0319397 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 9.435E-07 | 2.27E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.809E-06 | 5.785E-03 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 8.499E-06 | 6.816E-03 |
|---|
| interphase | GO:0051325 |  | 9.332E-06 | 5.613E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.057E-05 | 9.898E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.929E-05 | 0.01174514 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.131E-05 | 0.01763607 |
|---|
| fat cell differentiation | GO:0045444 |  | 6E-05 | 0.01804614 |
|---|
| base-excision repair | GO:0006284 |  | 6E-05 | 0.01604101 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.46E-05 | 0.01554366 |
|---|
| response to calcium ion | GO:0051592 |  | 1.839E-04 | 0.040224 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 9.435E-07 | 2.27E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 4.809E-06 | 5.785E-03 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 8.499E-06 | 6.816E-03 |
|---|
| interphase | GO:0051325 |  | 9.332E-06 | 5.613E-03 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.057E-05 | 9.898E-03 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 2.929E-05 | 0.01174514 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.131E-05 | 0.01763607 |
|---|
| fat cell differentiation | GO:0045444 |  | 6E-05 | 0.01804614 |
|---|
| base-excision repair | GO:0006284 |  | 6E-05 | 0.01604101 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.46E-05 | 0.01554366 |
|---|
| response to calcium ion | GO:0051592 |  | 1.839E-04 | 0.040224 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 8.963E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.172E-05 | 0.02432144 |
|---|
| interphase | GO:0051325 |  | 3.399E-05 | 0.01954254 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.02801271 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02334393 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| response to calcium ion | GO:0051592 |  | 3.919E-04 | 0.08193571 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 8.963E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.172E-05 | 0.02432144 |
|---|
| interphase | GO:0051325 |  | 3.399E-05 | 0.01954254 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.02801271 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02334393 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| response to calcium ion | GO:0051592 |  | 3.919E-04 | 0.08193571 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 8.963E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.172E-05 | 0.02432144 |
|---|
| interphase | GO:0051325 |  | 3.399E-05 | 0.01954254 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.02801271 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02334393 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| response to calcium ion | GO:0051592 |  | 3.919E-04 | 0.08193571 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| G1/S transition of mitotic cell cycle | GO:0000082 |  | 3.897E-06 | 8.963E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.07E-05 | 0.01230193 |
|---|
| interphase of mitotic cell cycle | GO:0051329 |  | 3.172E-05 | 0.02432144 |
|---|
| interphase | GO:0051325 |  | 3.399E-05 | 0.01954254 |
|---|
| cellular response to unfolded protein | GO:0034620 |  | 6.09E-05 | 0.02801271 |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 6.09E-05 | 0.02334393 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.063E-04 | 0.03494251 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.28E-04 | 0.03680434 |
|---|
| fat cell differentiation | GO:0045444 |  | 1.396E-04 | 0.03567408 |
|---|
| base-excision repair | GO:0006284 |  | 1.396E-04 | 0.03210667 |
|---|
| response to calcium ion | GO:0051592 |  | 3.919E-04 | 0.08193571 |
|---|



