Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 5908-7-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1744 | 1.515e-02 | 5.695e-03 | 8.086e-02 |
|---|
| IPC-NIBC-129 | 0.1769 | 6.696e-02 | 5.581e-02 | 3.111e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2270 | 5.043e-02 | 4.041e-02 | 4.390e-01 |
|---|
| Loi_GPL570 | 0.2499 | 8.764e-02 | 1.110e-01 | 3.450e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1832 | 8.644e-03 | 4.041e-03 | 1.123e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.3230 | 7.816e-02 | 3.968e-02 | 4.231e-01 |
|---|
| Schmidt | 0.2205 | 2.644e-02 | 2.806e-02 | 1.700e-01 |
|---|
| Sotiriou | 0.2667 | 8.342e-02 | 3.231e-02 | 5.103e-01 |
|---|
| Wang | 0.1949 | 5.220e-02 | 3.704e-02 | 1.862e-01 |
|---|
| Zhang | 0.2775 | 7.299e-03 | 3.581e-03 | 3.610e-02 |
|---|
Expression data for subnetwork 5908-7-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
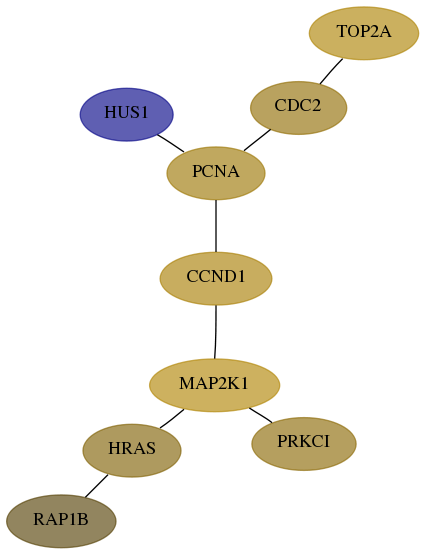
Score for each gene in subnetwork 5908-7-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| rap1b |   | 1 | 90 | 120 | 126 | 0.017 | 0.099 | 0.110 | 0.149 | 0.058 | 0.006 | -0.127 | 0.028 | 0.144 | 0.103 |
|---|
| pcna |   | 39 | 3 | 8 | 3 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.162 | 0.123 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| hras |   | 5 | 20 | 51 | 36 | 0.049 | -0.030 | 0.094 | 0.210 | 0.160 | 0.245 | 0.086 | 0.208 | 0.001 | 0.003 |
|---|
| top2a |   | 22 | 7 | 70 | 46 | 0.147 | 0.183 | 0.185 | 0.100 | 0.206 | 0.309 | 0.261 | 0.183 | 0.072 | 0.234 |
|---|
| hus1 |   | 6 | 17 | 22 | 17 | -0.058 | 0.197 | 0.116 | 0.109 | 0.101 | 0.160 | 0.072 | 0.082 | 0.081 | 0.071 |
|---|
| prkci |   | 1 | 90 | 120 | 126 | 0.065 | 0.104 | 0.012 | 0.170 | 0.112 | -0.170 | 0.196 | 0.128 | -0.054 | 0.124 |
|---|
| map2k1 |   | 3 | 39 | 51 | 42 | 0.154 | 0.096 | 0.093 | 0.013 | 0.095 | 0.207 | 0.082 | 0.096 | -0.043 | undef |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
GO Enrichment output for subnetwork 5908-7-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 1.924E-06 | 4.425E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 0.01021795 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 6.812E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 6.496E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 5.915E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.332E-05 | 5.106E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 4.377E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 3.83E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 4.763E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 1.864E-05 | 4.287E-03 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 5.193E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 2.42E-06 | 5.912E-03 |
|---|
| response to UV | GO:0009411 |  | 2.814E-06 | 3.437E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 2.972E-06 | 2.42E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 3.629E-06 | 2.216E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.629E-06 | 1.773E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.77E-06 | 2.756E-03 |
|---|
| DNA topological change | GO:0006265 |  | 6.77E-06 | 2.363E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 6.77E-06 | 2.067E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 6.77E-06 | 1.838E-03 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 6.77E-06 | 1.654E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 7.197E-06 | 1.598E-03 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.37E-06 | 8.109E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.37E-06 | 4.054E-03 |
|---|
| response to UV | GO:0009411 |  | 4.121E-06 | 3.305E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 4.871E-06 | 2.93E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 5.053E-06 | 2.432E-03 |
|---|
| DNA topological change | GO:0006265 |  | 9.426E-06 | 3.78E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 9.426E-06 | 3.24E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 9.426E-06 | 2.835E-03 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 9.426E-06 | 2.52E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.426E-06 | 2.268E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 1.178E-05 | 2.577E-03 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 2.42E-06 | 5.912E-03 |
|---|
| response to UV | GO:0009411 |  | 2.814E-06 | 3.437E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 2.972E-06 | 2.42E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 3.629E-06 | 2.216E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.629E-06 | 1.773E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.77E-06 | 2.756E-03 |
|---|
| DNA topological change | GO:0006265 |  | 6.77E-06 | 2.363E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 6.77E-06 | 2.067E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 6.77E-06 | 1.838E-03 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 6.77E-06 | 1.654E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 7.197E-06 | 1.598E-03 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.37E-06 | 8.109E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.37E-06 | 4.054E-03 |
|---|
| response to UV | GO:0009411 |  | 4.121E-06 | 3.305E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 4.871E-06 | 2.93E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 5.053E-06 | 2.432E-03 |
|---|
| DNA topological change | GO:0006265 |  | 9.426E-06 | 3.78E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 9.426E-06 | 3.24E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 9.426E-06 | 2.835E-03 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 9.426E-06 | 2.52E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.426E-06 | 2.268E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 1.178E-05 | 2.577E-03 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.37E-06 | 8.109E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.37E-06 | 4.054E-03 |
|---|
| response to UV | GO:0009411 |  | 4.121E-06 | 3.305E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 4.871E-06 | 2.93E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 5.053E-06 | 2.432E-03 |
|---|
| DNA topological change | GO:0006265 |  | 9.426E-06 | 3.78E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 9.426E-06 | 3.24E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 9.426E-06 | 2.835E-03 |
|---|
| regulation of synaptic transmission. GABAergic | GO:0032228 |  | 9.426E-06 | 2.52E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 9.426E-06 | 2.268E-03 |
|---|
| phosphoinositide-mediated signaling | GO:0048015 |  | 1.178E-05 | 2.577E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 1.924E-06 | 4.425E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 0.01021795 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 6.812E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 6.496E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 5.915E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.332E-05 | 5.106E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 4.377E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 3.83E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 4.763E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 1.864E-05 | 4.287E-03 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 5.193E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 1.924E-06 | 4.425E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 0.01021795 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 6.812E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 6.496E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 5.915E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.332E-05 | 5.106E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 4.377E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 3.83E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 4.763E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 1.864E-05 | 4.287E-03 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 5.193E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 1.924E-06 | 4.425E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 0.01021795 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 6.812E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 6.496E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 5.915E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.332E-05 | 5.106E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 4.377E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 3.83E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 4.763E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 1.864E-05 | 4.287E-03 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 5.193E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 1.924E-06 | 4.425E-03 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 0.01021795 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 6.812E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 6.496E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 5.915E-03 |
|---|
| photoreceptor cell development | GO:0042461 |  | 1.332E-05 | 5.106E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 4.377E-03 |
|---|
| regulation of Rac protein signal transduction | GO:0035020 |  | 1.332E-05 | 3.83E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.864E-05 | 4.763E-03 |
|---|
| establishment or maintenance of epithelial cell apical/basal polarity | GO:0045197 |  | 1.864E-05 | 4.287E-03 |
|---|
| positive regulation of small GTPase mediated signal transduction | GO:0051057 |  | 2.484E-05 | 5.193E-03 |
|---|



