Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 5424-6-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1704 | 1.724e-02 | 6.791e-03 | 8.947e-02 |
|---|
| IPC-NIBC-129 | 0.2445 | 6.638e-02 | 5.527e-02 | 3.088e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2207 | 1.328e-02 | 8.498e-03 | 1.595e-01 |
|---|
| Loi_GPL570 | 0.2506 | 9.917e-02 | 1.244e-01 | 3.770e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1667 | 1.011e-02 | 4.988e-03 | 1.347e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.2999 | 7.030e-02 | 3.464e-02 | 3.938e-01 |
|---|
| Schmidt | 0.2111 | 3.250e-02 | 3.434e-02 | 2.013e-01 |
|---|
| Sotiriou | 0.2536 | 1.087e-01 | 4.712e-02 | 5.950e-01 |
|---|
| Wang | 0.2478 | 2.083e-02 | 1.133e-02 | 9.051e-02 |
|---|
| Zhang | 0.2872 | 1.079e-02 | 5.544e-03 | 4.974e-02 |
|---|
Expression data for subnetwork 5424-6-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
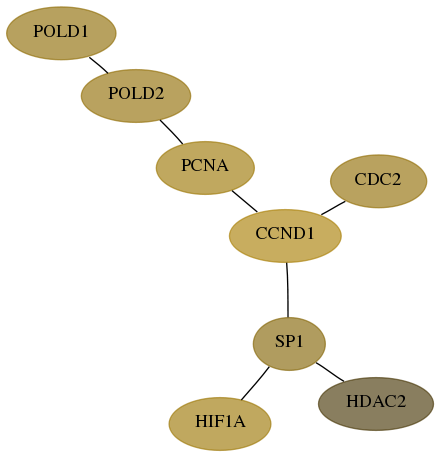
Score for each gene in subnetwork 5424-6-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| pold2 |   | 3 | 39 | 89 | 70 | 0.073 | 0.180 | 0.204 | 0.131 | 0.150 | 0.186 | 0.146 | 0.126 | -0.084 | 0.072 |
|---|
| pcna |   | 39 | 3 | 8 | 3 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.162 | 0.123 |
|---|
| hif1a |   | 7 | 15 | 16 | 13 | 0.097 | 0.246 | 0.089 | -0.009 | 0.101 | 0.011 | -0.043 | 0.119 | 0.172 | 0.129 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| sp1 |   | 31 | 5 | 16 | 8 | 0.054 | 0.186 | 0.147 | 0.242 | 0.125 | 0.202 | 0.284 | 0.227 | 0.194 | 0.091 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| hdac2 |   | 16 | 10 | 64 | 39 | 0.012 | 0.245 | 0.030 | 0.077 | 0.070 | 0.183 | 0.222 | -0.058 | 0.086 | 0.133 |
|---|
| pold1 |   | 2 | 53 | 89 | 81 | 0.069 | 0.107 | 0.160 | -0.033 | 0.102 | 0.121 | 0.144 | 0.148 | -0.048 | 0.006 |
|---|
GO Enrichment output for subnetwork 5424-6-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.328E-10 | 7.655E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 4.878E-08 | 5.61E-05 |
|---|
| base-excision repair | GO:0006284 |  | 6.443E-07 | 4.94E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 2.574E-03 |
|---|
| response to UV | GO:0009411 |  | 4.787E-06 | 2.202E-03 |
|---|
| regulation of hormone biosynthetic process | GO:0046885 |  | 5.098E-06 | 1.954E-03 |
|---|
| embryonic hemopoiesis | GO:0035162 |  | 5.098E-06 | 1.675E-03 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 5.098E-06 | 1.466E-03 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 5.098E-06 | 1.173E-03 |
|---|
| positive regulation of nitric-oxide synthase activity | GO:0051000 |  | 7.644E-06 | 1.598E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.732E-11 | 6.675E-08 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 5.876E-09 | 7.177E-06 |
|---|
| base-excision repair | GO:0006284 |  | 1.584E-07 | 1.29E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.325E-07 | 5.085E-04 |
|---|
| response to UV | GO:0009411 |  | 1.261E-06 | 6.16E-04 |
|---|
| hemoglobin biosynthetic process | GO:0042541 |  | 1.44E-06 | 5.862E-04 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 1.44E-06 | 5.024E-04 |
|---|
| mRNA transcription | GO:0009299 |  | 1.44E-06 | 4.396E-04 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 1.44E-06 | 3.908E-04 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 1.44E-06 | 3.517E-04 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 1.44E-06 | 3.197E-04 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.299E-11 | 1.275E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 1.058E-08 | 1.273E-05 |
|---|
| base-excision repair | GO:0006284 |  | 2.323E-07 | 1.863E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.366E-06 | 8.216E-04 |
|---|
| response to UV | GO:0009411 |  | 1.848E-06 | 8.891E-04 |
|---|
| hemoglobin biosynthetic process | GO:0042541 |  | 2.005E-06 | 8.04E-04 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 2.005E-06 | 6.892E-04 |
|---|
| mRNA transcription | GO:0009299 |  | 2.005E-06 | 6.03E-04 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 2.005E-06 | 5.36E-04 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.005E-06 | 4.824E-04 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 2.005E-06 | 4.386E-04 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 2.732E-11 | 6.675E-08 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 5.876E-09 | 7.177E-06 |
|---|
| base-excision repair | GO:0006284 |  | 1.584E-07 | 1.29E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.325E-07 | 5.085E-04 |
|---|
| response to UV | GO:0009411 |  | 1.261E-06 | 6.16E-04 |
|---|
| hemoglobin biosynthetic process | GO:0042541 |  | 1.44E-06 | 5.862E-04 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 1.44E-06 | 5.024E-04 |
|---|
| mRNA transcription | GO:0009299 |  | 1.44E-06 | 4.396E-04 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 1.44E-06 | 3.908E-04 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 1.44E-06 | 3.517E-04 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 1.44E-06 | 3.197E-04 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.299E-11 | 1.275E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 1.058E-08 | 1.273E-05 |
|---|
| base-excision repair | GO:0006284 |  | 2.323E-07 | 1.863E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.366E-06 | 8.216E-04 |
|---|
| response to UV | GO:0009411 |  | 1.848E-06 | 8.891E-04 |
|---|
| hemoglobin biosynthetic process | GO:0042541 |  | 2.005E-06 | 8.04E-04 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 2.005E-06 | 6.892E-04 |
|---|
| mRNA transcription | GO:0009299 |  | 2.005E-06 | 6.03E-04 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 2.005E-06 | 5.36E-04 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.005E-06 | 4.824E-04 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 2.005E-06 | 4.386E-04 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 5.299E-11 | 1.275E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 1.058E-08 | 1.273E-05 |
|---|
| base-excision repair | GO:0006284 |  | 2.323E-07 | 1.863E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.366E-06 | 8.216E-04 |
|---|
| response to UV | GO:0009411 |  | 1.848E-06 | 8.891E-04 |
|---|
| hemoglobin biosynthetic process | GO:0042541 |  | 2.005E-06 | 8.04E-04 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 2.005E-06 | 6.892E-04 |
|---|
| mRNA transcription | GO:0009299 |  | 2.005E-06 | 6.03E-04 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 2.005E-06 | 5.36E-04 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.005E-06 | 4.824E-04 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 2.005E-06 | 4.386E-04 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.328E-10 | 7.655E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 4.878E-08 | 5.61E-05 |
|---|
| base-excision repair | GO:0006284 |  | 6.443E-07 | 4.94E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 2.574E-03 |
|---|
| response to UV | GO:0009411 |  | 4.787E-06 | 2.202E-03 |
|---|
| regulation of hormone biosynthetic process | GO:0046885 |  | 5.098E-06 | 1.954E-03 |
|---|
| embryonic hemopoiesis | GO:0035162 |  | 5.098E-06 | 1.675E-03 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 5.098E-06 | 1.466E-03 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 5.098E-06 | 1.173E-03 |
|---|
| positive regulation of nitric-oxide synthase activity | GO:0051000 |  | 7.644E-06 | 1.598E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.328E-10 | 7.655E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 4.878E-08 | 5.61E-05 |
|---|
| base-excision repair | GO:0006284 |  | 6.443E-07 | 4.94E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 2.574E-03 |
|---|
| response to UV | GO:0009411 |  | 4.787E-06 | 2.202E-03 |
|---|
| regulation of hormone biosynthetic process | GO:0046885 |  | 5.098E-06 | 1.954E-03 |
|---|
| embryonic hemopoiesis | GO:0035162 |  | 5.098E-06 | 1.675E-03 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 5.098E-06 | 1.466E-03 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 5.098E-06 | 1.173E-03 |
|---|
| positive regulation of nitric-oxide synthase activity | GO:0051000 |  | 7.644E-06 | 1.598E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.328E-10 | 7.655E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 4.878E-08 | 5.61E-05 |
|---|
| base-excision repair | GO:0006284 |  | 6.443E-07 | 4.94E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 2.574E-03 |
|---|
| response to UV | GO:0009411 |  | 4.787E-06 | 2.202E-03 |
|---|
| regulation of hormone biosynthetic process | GO:0046885 |  | 5.098E-06 | 1.954E-03 |
|---|
| embryonic hemopoiesis | GO:0035162 |  | 5.098E-06 | 1.675E-03 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 5.098E-06 | 1.466E-03 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 5.098E-06 | 1.173E-03 |
|---|
| positive regulation of nitric-oxide synthase activity | GO:0051000 |  | 7.644E-06 | 1.598E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| nucleotide-excision repair. DNA gap filling | GO:0006297 |  | 3.328E-10 | 7.655E-07 |
|---|
| nucleotide-excision repair | GO:0006289 |  | 4.878E-08 | 5.61E-05 |
|---|
| base-excision repair | GO:0006284 |  | 6.443E-07 | 4.94E-04 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 4.477E-06 | 2.574E-03 |
|---|
| response to UV | GO:0009411 |  | 4.787E-06 | 2.202E-03 |
|---|
| regulation of hormone biosynthetic process | GO:0046885 |  | 5.098E-06 | 1.954E-03 |
|---|
| embryonic hemopoiesis | GO:0035162 |  | 5.098E-06 | 1.675E-03 |
|---|
| positive regulation of glycolysis | GO:0045821 |  | 5.098E-06 | 1.466E-03 |
|---|
| maintenance of fidelity during DNA-dependent DNA replication | GO:0045005 |  | 5.098E-06 | 1.303E-03 |
|---|
| positive regulation of chemokine production | GO:0032722 |  | 5.098E-06 | 1.173E-03 |
|---|
| positive regulation of nitric-oxide synthase activity | GO:0051000 |  | 7.644E-06 | 1.598E-03 |
|---|



