Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 1431-1-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.1668 | 1.740e-02 | 6.878e-03 | 9.012e-02 |
|---|
| IPC-NIBC-129 | 0.1787 | 4.507e-02 | 3.555e-02 | 2.174e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2203 | 4.868e-02 | 3.879e-02 | 4.290e-01 |
|---|
| Loi_GPL570 | 0.2811 | 1.103e-01 | 1.372e-01 | 4.063e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1857 | 1.166e-02 | 6.035e-03 | 1.581e-01 |
|---|
| Pawitan_GPL96-GPL97 | 0.3179 | 7.432e-02 | 3.720e-02 | 4.090e-01 |
|---|
| Schmidt | 0.2030 | 4.117e-02 | 4.327e-02 | 2.430e-01 |
|---|
| Sotiriou | 0.2386 | 8.015e-02 | 3.051e-02 | 4.979e-01 |
|---|
| Wang | 0.2264 | 3.026e-02 | 1.839e-02 | 1.219e-01 |
|---|
| Zhang | 0.2821 | 7.962e-03 | 3.946e-03 | 3.878e-02 |
|---|
Expression data for subnetwork 1431-1-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
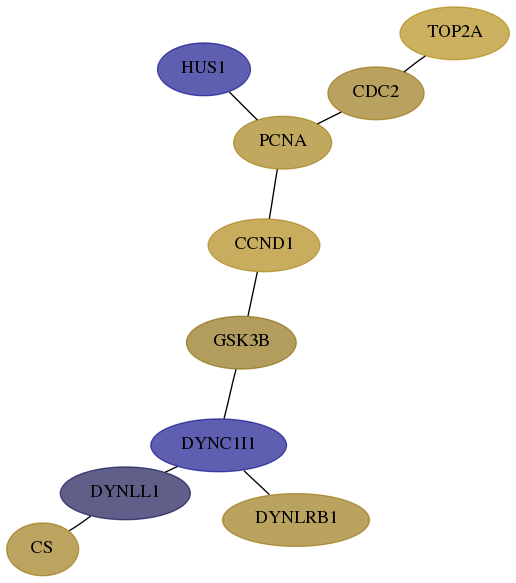
Score for each gene in subnetwork 1431-1-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| pcna |   | 39 | 3 | 8 | 3 | 0.096 | 0.202 | 0.109 | 0.180 | 0.118 | 0.256 | 0.151 | 0.059 | 0.162 | 0.123 |
|---|
| hus1 |   | 6 | 17 | 22 | 17 | -0.058 | 0.197 | 0.116 | 0.109 | 0.101 | 0.160 | 0.072 | 0.082 | 0.081 | 0.071 |
|---|
| gsk3b |   | 2 | 53 | 111 | 101 | 0.057 | 0.003 | 0.058 | 0.029 | 0.154 | -0.068 | 0.119 | 0.070 | 0.061 | 0.146 |
|---|
| dynlrb1 |   | 1 | 90 | 111 | 118 | 0.077 | 0.032 | 0.076 | 0.289 | 0.148 | 0.218 | -0.037 | 0.091 | 0.042 | 0.019 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| cs |   | 1 | 90 | 111 | 118 | 0.079 | 0.125 | 0.055 | 0.121 | 0.097 | 0.010 | -0.052 | 0.067 | 0.051 | -0.157 |
|---|
| dync1i1 |   | 3 | 39 | 45 | 38 | -0.058 | 0.102 | 0.055 | 0.124 | 0.054 | 0.056 | 0.082 | 0.036 | 0.119 | 0.166 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| top2a |   | 22 | 7 | 70 | 46 | 0.147 | 0.183 | 0.185 | 0.100 | 0.206 | 0.309 | 0.261 | 0.183 | 0.072 | 0.234 |
|---|
| dynll1 |   | 2 | 53 | 45 | 43 | -0.012 | 0.108 | 0.088 | 0.087 | 0.098 | 0.128 | 0.113 | 0.093 | 0.158 | 0.074 |
|---|
GO Enrichment output for subnetwork 1431-1-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 2.64E-08 | 6.072E-05 |
|---|
| visual behavior | GO:0007632 |  | 1.005E-06 | 1.156E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.005E-06 | 7.704E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.336E-06 | 7.685E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 4.087E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 3.406E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 3.712E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 3.697E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 1.332E-05 | 3.404E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.332E-05 | 3.064E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 2.785E-03 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| visual behavior | GO:0007632 |  | 3.615E-07 | 8.832E-04 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.301E-07 | 6.475E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 7.441E-07 | 6.06E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.653E-06 | 2.231E-03 |
|---|
| response to UV | GO:0009411 |  | 5.287E-06 | 2.583E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 5.477E-06 | 2.23E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 5.584E-06 | 1.949E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 7.665E-06 | 2.341E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.022E-05 | 2.773E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.022E-05 | 2.496E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.022E-05 | 2.269E-03 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| visual behavior | GO:0007632 |  | 3.74E-07 | 8.998E-04 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 4.853E-07 | 5.838E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.904E-07 | 5.537E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.766E-06 | 2.265E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.766E-06 | 1.812E-03 |
|---|
| response to UV | GO:0009411 |  | 4.888E-06 | 1.96E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 5.777E-06 | 1.986E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 7.903E-06 | 2.377E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.053E-05 | 2.816E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.053E-05 | 2.534E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.053E-05 | 2.304E-03 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| visual behavior | GO:0007632 |  | 3.615E-07 | 8.832E-04 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 5.301E-07 | 6.475E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 7.441E-07 | 6.06E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.653E-06 | 2.231E-03 |
|---|
| response to UV | GO:0009411 |  | 5.287E-06 | 2.583E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 5.477E-06 | 2.23E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 5.584E-06 | 1.949E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 7.665E-06 | 2.341E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.022E-05 | 2.773E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.022E-05 | 2.496E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.022E-05 | 2.269E-03 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| visual behavior | GO:0007632 |  | 3.74E-07 | 8.998E-04 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 4.853E-07 | 5.838E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.904E-07 | 5.537E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.766E-06 | 2.265E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.766E-06 | 1.812E-03 |
|---|
| response to UV | GO:0009411 |  | 4.888E-06 | 1.96E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 5.777E-06 | 1.986E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 7.903E-06 | 2.377E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.053E-05 | 2.816E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.053E-05 | 2.534E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.053E-05 | 2.304E-03 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| visual behavior | GO:0007632 |  | 3.74E-07 | 8.998E-04 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 4.853E-07 | 5.838E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 6.904E-07 | 5.537E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 3.766E-06 | 2.265E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 3.766E-06 | 1.812E-03 |
|---|
| response to UV | GO:0009411 |  | 4.888E-06 | 1.96E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 5.777E-06 | 1.986E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 7.903E-06 | 2.377E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.053E-05 | 2.816E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.053E-05 | 2.534E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.053E-05 | 2.304E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 2.64E-08 | 6.072E-05 |
|---|
| visual behavior | GO:0007632 |  | 1.005E-06 | 1.156E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.005E-06 | 7.704E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.336E-06 | 7.685E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 4.087E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 3.406E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 3.712E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 3.697E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 1.332E-05 | 3.404E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.332E-05 | 3.064E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 2.785E-03 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 2.64E-08 | 6.072E-05 |
|---|
| visual behavior | GO:0007632 |  | 1.005E-06 | 1.156E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.005E-06 | 7.704E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.336E-06 | 7.685E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 4.087E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 3.406E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 3.712E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 3.697E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 1.332E-05 | 3.404E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.332E-05 | 3.064E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 2.785E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 2.64E-08 | 6.072E-05 |
|---|
| visual behavior | GO:0007632 |  | 1.005E-06 | 1.156E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.005E-06 | 7.704E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.336E-06 | 7.685E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 4.087E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 3.406E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 3.712E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 3.697E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 1.332E-05 | 3.404E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.332E-05 | 3.064E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 2.785E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to light stimulus | GO:0009416 |  | 2.64E-08 | 6.072E-05 |
|---|
| visual behavior | GO:0007632 |  | 1.005E-06 | 1.156E-03 |
|---|
| response to endoplasmic reticulum stress | GO:0034976 |  | 1.005E-06 | 7.704E-04 |
|---|
| ER-nuclear signaling pathway | GO:0006984 |  | 1.336E-06 | 7.685E-04 |
|---|
| positive regulation of viral genome replication | GO:0045070 |  | 8.885E-06 | 4.087E-03 |
|---|
| regulation of retroviral genome replication | GO:0045091 |  | 8.885E-06 | 3.406E-03 |
|---|
| response to UV | GO:0009411 |  | 1.13E-05 | 3.712E-03 |
|---|
| regulation of DNA replication | GO:0006275 |  | 1.286E-05 | 3.697E-03 |
|---|
| regulation of protein export from nucleus | GO:0046825 |  | 1.332E-05 | 3.404E-03 |
|---|
| ER overload response | GO:0006983 |  | 1.332E-05 | 3.064E-03 |
|---|
| DNA topological change | GO:0006265 |  | 1.332E-05 | 2.785E-03 |
|---|



