Study run-b1
Study informations
127 subnetworks in total page | file
236 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 1294-1-real
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 |
|---|
| Desmedt | 0.2025 | 5.936e-03 | 1.581e-03 | 3.828e-02 |
|---|
| IPC-NIBC-129 | 0.2305 | 4.523e-02 | 3.570e-02 | 2.181e-01 |
|---|
| Ivshina_GPL96-GPL97 | 0.2716 | 1.755e-02 | 1.179e-02 | 2.016e-01 |
|---|
| Loi_GPL570 | 0.2808 | 1.139e-01 | 1.412e-01 | 4.154e-01 |
|---|
| Loi_GPL96-GPL97 | 0.1735 | 2.790e-03 | 8.720e-04 | 2.566e-02 |
|---|
| Pawitan_GPL96-GPL97 | 0.3333 | 4.419e-01 | 3.639e-01 | 9.397e-01 |
|---|
| Schmidt | 0.2005 | 9.798e-03 | 1.064e-02 | 7.116e-02 |
|---|
| Sotiriou | 0.3291 | 9.753e-02 | 4.038e-02 | 5.599e-01 |
|---|
| Wang | 0.0805 | 3.491e-01 | 3.784e-01 | 6.788e-01 |
|---|
| Zhang | 0.1105 | 6.135e-03 | 2.950e-03 | 3.128e-02 |
|---|
Expression data for subnetwork 1294-1-real in each dataset
Desmedt |
IPC-NIBC-129 |
Ivshina_GPL96-GPL97 |
Loi_GPL570 |
Loi_GPL96-GPL97 |
Pawitan_GPL96-GPL97 |
Schmidt |
Sotiriou |
Wang |
Zhang |
Subnetwork structure for each dataset
- Desmedt
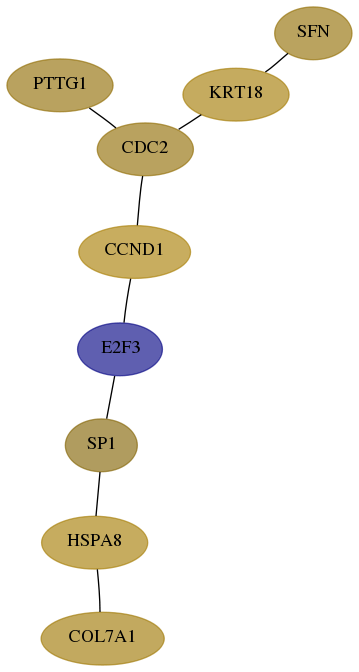
Score for each gene in subnetwork 1294-1-real in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC-NIBC-129 | Ivshina_GPL96-GPL97 | Loi_GPL570 | Loi_GPL96-GPL97 | Pawitan_GPL96-GPL97 | Schmidt | Sotiriou | Wang | Zhang |
|---|
| pttg1 |   | 4 | 27 | 173 | 150 | 0.072 | 0.270 | 0.175 | 0.183 | 0.200 | 0.334 | 0.224 | 0.204 | 0.147 | 0.210 |
|---|
| krt18 |   | 13 | 12 | 99 | 67 | 0.112 | -0.078 | 0.147 | 0.150 | 0.161 | 0.083 | -0.099 | 0.118 | 0.086 | -0.114 |
|---|
| col7a1 |   | 1 | 90 | 186 | 189 | 0.103 | 0.095 | 0.098 | 0.011 | 0.070 | -0.002 | 0.092 | 0.035 | 0.056 | 0.028 |
|---|
| ccnd1 |   | 97 | 2 | 8 | 2 | 0.127 | -0.183 | 0.062 | 0.190 | 0.157 | 0.105 | -0.107 | 0.174 | 0.106 | 0.144 |
|---|
| sp1 |   | 31 | 5 | 16 | 8 | 0.054 | 0.186 | 0.147 | 0.242 | 0.125 | 0.202 | 0.284 | 0.227 | 0.194 | 0.091 |
|---|
| hspa8 |   | 1 | 90 | 186 | 189 | 0.122 | 0.064 | 0.054 | -0.044 | 0.048 | 0.077 | -0.171 | 0.088 | -0.094 | -0.140 |
|---|
| e2f3 |   | 3 | 39 | 87 | 68 | -0.054 | 0.341 | 0.099 | 0.140 | 0.072 | 0.097 | 0.340 | 0.081 | 0.017 | 0.032 |
|---|
| cdc2 |   | 99 | 1 | 1 | 1 | 0.073 | 0.247 | 0.206 | 0.172 | 0.247 | 0.282 | 0.226 | 0.221 | 0.067 | 0.297 |
|---|
| sfn |   | 4 | 27 | 1 | 5 | 0.077 | 0.137 | 0.111 | 0.172 | 0.112 | 0.219 | 0.100 | 0.147 | 0.025 | 0.079 |
|---|
GO Enrichment output for subnetwork 1294-1-real in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 5.717E-09 | 1.315E-05 |
|---|
| response to protein stimulus | GO:0051789 |  | 7.907E-09 | 9.093E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.719E-06 | 6.684E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.275E-05 | 7.333E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.645E-05 | 7.566E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.786E-05 | 6.845E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 2.192E-05 | 7.202E-03 |
|---|
| positive regulation of cell growth | GO:0030307 |  | 5.156E-05 | 0.01482233 |
|---|
| negative regulation of caspase activity | GO:0043154 |  | 7.101E-05 | 0.01814669 |
|---|
| release of cytochrome c from mitochondria | GO:0001836 |  | 7.101E-05 | 0.01633203 |
|---|
| positive regulation of cell size | GO:0045793 |  | 8.189E-05 | 0.01712235 |
|---|
IPC-NIBC-129 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 5.618E-10 | 1.372E-06 |
|---|
| response to protein stimulus | GO:0051789 |  | 7.8E-10 | 9.528E-07 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.551E-06 | 1.263E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.151E-06 | 1.314E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.617E-06 | 1.278E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.226E-06 | 1.313E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 4.271E-06 | 1.491E-03 |
|---|
| tumor necrosis factor-mediated signaling pathway | GO:0033209 |  | 4.515E-06 | 1.379E-03 |
|---|
| Golgi to plasma membrane transport | GO:0006893 |  | 4.515E-06 | 1.226E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.018E-06 | 1.47E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 1.181E-05 | 2.623E-03 |
|---|
Ivshina_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 9.576E-10 | 2.304E-06 |
|---|
| response to protein stimulus | GO:0051789 |  | 1.355E-09 | 1.63E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.543E-06 | 2.04E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.996E-06 | 1.802E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.289E-06 | 2.064E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 4.492E-06 | 1.801E-03 |
|---|
| tumor necrosis factor-mediated signaling pathway | GO:0033209 |  | 6.287E-06 | 2.161E-03 |
|---|
| Golgi to plasma membrane transport | GO:0006893 |  | 6.287E-06 | 1.891E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 6.998E-06 | 1.871E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 8.38E-06 | 2.016E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 1.077E-05 | 2.356E-03 |
|---|
Loi_GPL570 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 5.618E-10 | 1.372E-06 |
|---|
| response to protein stimulus | GO:0051789 |  | 7.8E-10 | 9.528E-07 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 1.551E-06 | 1.263E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.151E-06 | 1.314E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 2.617E-06 | 1.278E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 3.226E-06 | 1.313E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 4.271E-06 | 1.491E-03 |
|---|
| tumor necrosis factor-mediated signaling pathway | GO:0033209 |  | 4.515E-06 | 1.379E-03 |
|---|
| Golgi to plasma membrane transport | GO:0006893 |  | 4.515E-06 | 1.226E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 6.018E-06 | 1.47E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 1.181E-05 | 2.623E-03 |
|---|
Loi_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 9.576E-10 | 2.304E-06 |
|---|
| response to protein stimulus | GO:0051789 |  | 1.355E-09 | 1.63E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.543E-06 | 2.04E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.996E-06 | 1.802E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.289E-06 | 2.064E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 4.492E-06 | 1.801E-03 |
|---|
| tumor necrosis factor-mediated signaling pathway | GO:0033209 |  | 6.287E-06 | 2.161E-03 |
|---|
| Golgi to plasma membrane transport | GO:0006893 |  | 6.287E-06 | 1.891E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 6.998E-06 | 1.871E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 8.38E-06 | 2.016E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 1.077E-05 | 2.356E-03 |
|---|
Pawitan_GPL96-GPL97 file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 9.576E-10 | 2.304E-06 |
|---|
| response to protein stimulus | GO:0051789 |  | 1.355E-09 | 1.63E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 2.543E-06 | 2.04E-03 |
|---|
| meiotic chromosome segregation | GO:0045132 |  | 2.996E-06 | 1.802E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 4.289E-06 | 2.064E-03 |
|---|
| establishment of spindle localization | GO:0051293 |  | 4.492E-06 | 1.801E-03 |
|---|
| tumor necrosis factor-mediated signaling pathway | GO:0033209 |  | 6.287E-06 | 2.161E-03 |
|---|
| Golgi to plasma membrane transport | GO:0006893 |  | 6.287E-06 | 1.891E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 6.998E-06 | 1.871E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 8.38E-06 | 2.016E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 1.077E-05 | 2.356E-03 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 5.717E-09 | 1.315E-05 |
|---|
| response to protein stimulus | GO:0051789 |  | 7.907E-09 | 9.093E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.719E-06 | 6.684E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.275E-05 | 7.333E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.645E-05 | 7.566E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.786E-05 | 6.845E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 2.192E-05 | 7.202E-03 |
|---|
| positive regulation of cell growth | GO:0030307 |  | 5.156E-05 | 0.01482233 |
|---|
| negative regulation of caspase activity | GO:0043154 |  | 7.101E-05 | 0.01814669 |
|---|
| release of cytochrome c from mitochondria | GO:0001836 |  | 7.101E-05 | 0.01633203 |
|---|
| positive regulation of cell size | GO:0045793 |  | 8.189E-05 | 0.01712235 |
|---|
Sotiriou file
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 5.717E-09 | 1.315E-05 |
|---|
| response to protein stimulus | GO:0051789 |  | 7.907E-09 | 9.093E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.719E-06 | 6.684E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.275E-05 | 7.333E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.645E-05 | 7.566E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.786E-05 | 6.845E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 2.192E-05 | 7.202E-03 |
|---|
| positive regulation of cell growth | GO:0030307 |  | 5.156E-05 | 0.01482233 |
|---|
| negative regulation of caspase activity | GO:0043154 |  | 7.101E-05 | 0.01814669 |
|---|
| release of cytochrome c from mitochondria | GO:0001836 |  | 7.101E-05 | 0.01633203 |
|---|
| positive regulation of cell size | GO:0045793 |  | 8.189E-05 | 0.01712235 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 5.717E-09 | 1.315E-05 |
|---|
| response to protein stimulus | GO:0051789 |  | 7.907E-09 | 9.093E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.719E-06 | 6.684E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.275E-05 | 7.333E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.645E-05 | 7.566E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.786E-05 | 6.845E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 2.192E-05 | 7.202E-03 |
|---|
| positive regulation of cell growth | GO:0030307 |  | 5.156E-05 | 0.01482233 |
|---|
| negative regulation of caspase activity | GO:0043154 |  | 7.101E-05 | 0.01814669 |
|---|
| release of cytochrome c from mitochondria | GO:0001836 |  | 7.101E-05 | 0.01633203 |
|---|
| positive regulation of cell size | GO:0045793 |  | 8.189E-05 | 0.01712235 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| response to unfolded protein | GO:0006986 |  | 5.717E-09 | 1.315E-05 |
|---|
| response to protein stimulus | GO:0051789 |  | 7.907E-09 | 9.093E-06 |
|---|
| positive regulation of cell cycle | GO:0045787 |  | 8.719E-06 | 6.684E-03 |
|---|
| regulation of cyclin-dependent protein kinase activity | GO:0000079 |  | 1.275E-05 | 7.333E-03 |
|---|
| positive regulation of cyclin-dependent protein kinase activity | GO:0045737 |  | 1.645E-05 | 7.566E-03 |
|---|
| chromosome segregation | GO:0007059 |  | 1.786E-05 | 6.845E-03 |
|---|
| keratinocyte proliferation | GO:0043616 |  | 2.192E-05 | 7.202E-03 |
|---|
| positive regulation of cell growth | GO:0030307 |  | 5.156E-05 | 0.01482233 |
|---|
| negative regulation of caspase activity | GO:0043154 |  | 7.101E-05 | 0.01814669 |
|---|
| release of cytochrome c from mitochondria | GO:0001836 |  | 7.101E-05 | 0.01633203 |
|---|
| positive regulation of cell size | GO:0045793 |  | 8.189E-05 | 0.01712235 |
|---|



