Study VanDeVijver-ER-pos
Study informations
14 subnetworks in total page | file
272 genes associated page | file
Enriched GO terms page
General informations
General Index page
Study Index page
Subnetwork 119
score
| Dataset | Score | P-val 1 | P-val 2 | P-val 3 | Fisher Score |
|---|
| Desmedt | 0.5287 | 0.000e+00 | 0.000e+00 | 2.420e-04 | 0.000e+00 |
|---|
| IPC | 0.5688 | 2.609e-02 | 1.520e-02 | 9.449e-01 | 3.749e-04 |
|---|
| Loi | 0.4921 | 0.000e+00 | 1.000e-06 | 3.739e-03 | 0.000e+00 |
|---|
| Schmidt | 0.2849 | 1.836e-02 | 5.594e-03 | 3.411e-01 | 3.503e-05 |
|---|
| Wang | 0.3415 | 1.070e-04 | 2.000e-06 | 4.491e-02 | 9.610e-12 |
|---|
Expression data for subnetwork 119 in each dataset
Desmedt |
IPC |
Loi |
Schmidt |
Wang |
Subnetwork structure for each dataset
- Desmedt
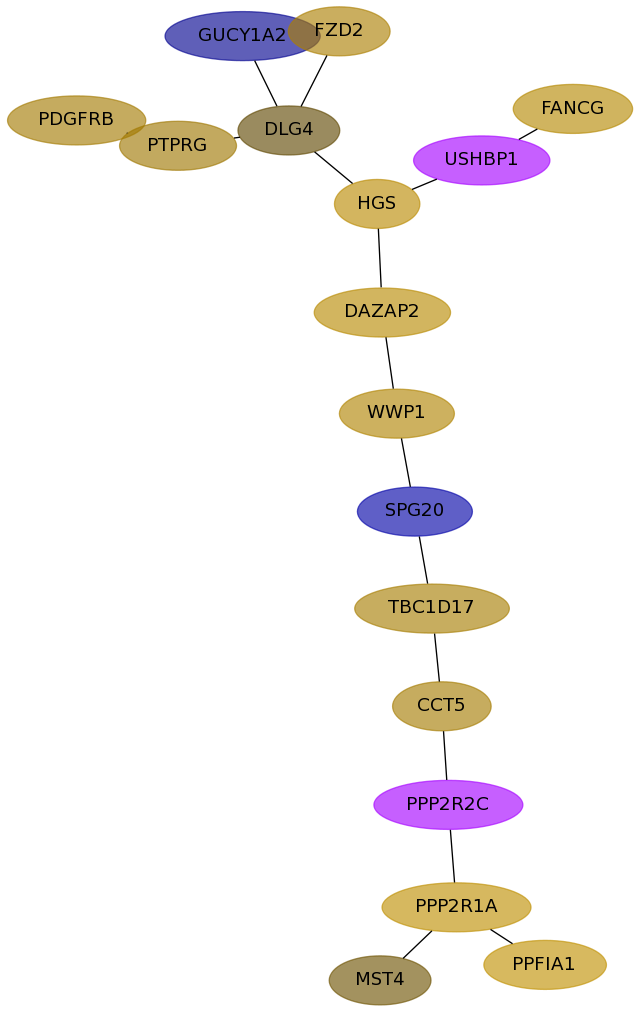
Score for each gene in subnetwork 119 in each dataset
| Gene Symbol | Links | Frequency | Frequency Rank | Subnetwork score rank | Global rank |
Desmedt | IPC | Loi | Schmidt | Wang |
|---|
| DLG4 |   | 1 | 59 | 65 | 72 | 0.023 | -0.178 | -0.047 | 0.146 | undef |
|---|
| FANCG |   | 1 | 59 | 65 | 72 | 0.174 | 0.131 | 0.179 | 0.104 | 0.155 |
|---|
| GUCY1A2 |   | 1 | 59 | 65 | 72 | -0.071 | 0.134 | 0.016 | 0.008 | -0.061 |
|---|
| FZD2 |   | 1 | 59 | 65 | 72 | 0.137 | -0.062 | 0.160 | 0.076 | 0.078 |
|---|
| PPFIA1 |   | 1 | 59 | 65 | 72 | 0.226 | 0.129 | 0.107 | 0.064 | 0.208 |
|---|
| DAZAP2 |   | 2 | 27 | 65 | 65 | 0.186 | -0.174 | 0.074 | -0.249 | -0.046 |
|---|
| PTPRG |   | 1 | 59 | 65 | 72 | 0.104 | -0.029 | 0.024 | -0.125 | -0.026 |
|---|
| CCT5 |   | 2 | 27 | 65 | 65 | 0.122 | 0.132 | 0.196 | 0.122 | 0.139 |
|---|
| TBC1D17 |   | 2 | 27 | 65 | 65 | 0.126 | -0.165 | 0.100 | -0.037 | -0.031 |
|---|
| HGS |   | 1 | 59 | 65 | 72 | 0.194 | 0.010 | 0.109 | 0.071 | 0.004 |
|---|
| PPP2R1A |   | 2 | 27 | 65 | 65 | 0.215 | -0.149 | 0.172 | -0.047 | -0.085 |
|---|
| MST4 |   | 2 | 27 | 65 | 65 | 0.033 | 0.069 | 0.222 | 0.226 | 0.070 |
|---|
| WWP1 |   | 2 | 27 | 65 | 65 | 0.154 | -0.081 | 0.132 | 0.007 | 0.028 |
|---|
| PPP2R2C |   | 2 | 27 | 65 | 65 | undef | -0.085 | 0.177 | undef | undef |
|---|
| SPG20 |   | 2 | 27 | 36 | 41 | -0.125 | -0.029 | 0.009 | -0.154 | 0.042 |
|---|
| PDGFRB |   | 3 | 13 | 1 | 5 | 0.116 | -0.093 | -0.038 | -0.159 | 0.110 |
|---|
| USHBP1 |   | 1 | 59 | 65 | 72 | undef | -0.055 | 0.005 | undef | undef |
|---|
GO Enrichment output for subnetwork 119 in each dataset
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| negative regulation of JAK-STAT cascade | GO:0046426 |  | 6.034E-08 | 1.388E-04 |
|---|
| negative regulation of protein kinase cascade | GO:0010741 |  | 4.414E-06 | 5.076E-03 |
|---|
| negative regulation of peptidyl-tyrosine phosphorylation | GO:0050732 |  | 1.508E-05 | 0.01156371 |
|---|
| regulation of JAK-STAT cascade | GO:0046425 |  | 1.924E-05 | 0.01106041 |
|---|
| establishment of tissue polarity | GO:0007164 |  | 2.261E-05 | 0.0103994 |
|---|
| synaptic vesicle endocytosis | GO:0048488 |  | 3.163E-05 | 0.01212338 |
|---|
| regulation of peptidyl-tyrosine phosphorylation | GO:0050730 |  | 6.541E-05 | 0.02149151 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 6.762E-05 | 0.01943946 |
|---|
| platelet-derived growth factor receptor signaling pathway | GO:0048008 |  | 6.762E-05 | 0.01727952 |
|---|
| entry into host cell | GO:0030260 |  | 6.762E-05 | 0.01555157 |
|---|
| ER-associated protein catabolic process | GO:0030433 |  | 8.258E-05 | 0.01726635 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| negative regulation of JAK-STAT cascade | GO:0046426 |  | 3.624E-08 | 8.852E-05 |
|---|
| negative regulation of protein kinase cascade | GO:0010741 |  | 4.134E-06 | 5.05E-03 |
|---|
| negative regulation of peptidyl-tyrosine phosphorylation | GO:0050732 |  | 1.052E-05 | 8.569E-03 |
|---|
| retrograde protein transport. ER to cytosol | GO:0030970 |  | 1.052E-05 | 6.427E-03 |
|---|
| regulation of JAK-STAT cascade | GO:0046425 |  | 1.739E-05 | 8.496E-03 |
|---|
| establishment of tissue polarity | GO:0007164 |  | 2.207E-05 | 8.986E-03 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 4.72E-05 | 0.01647178 |
|---|
| synaptic vesicle endocytosis | GO:0048488 |  | 4.72E-05 | 0.01441281 |
|---|
| regulation of peptidyl-tyrosine phosphorylation | GO:0050730 |  | 5.188E-05 | 0.0140837 |
|---|
| entry into host cell | GO:0030260 |  | 6.913E-05 | 0.01688884 |
|---|
| cGMP metabolic process | GO:0046068 |  | 8.165E-05 | 0.01813314 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| negative regulation of JAK-STAT cascade | GO:0046426 |  | 5.016E-08 | 1.207E-04 |
|---|
| negative regulation of protein kinase cascade | GO:0010741 |  | 5.713E-06 | 6.873E-03 |
|---|
| negative regulation of peptidyl-tyrosine phosphorylation | GO:0050732 |  | 1.309E-05 | 0.0104993 |
|---|
| retrograde protein transport. ER to cytosol | GO:0030970 |  | 1.309E-05 | 7.874E-03 |
|---|
| regulation of JAK-STAT cascade | GO:0046425 |  | 1.973E-05 | 9.494E-03 |
|---|
| establishment of tissue polarity | GO:0007164 |  | 2.745E-05 | 0.01100817 |
|---|
| synaptic vesicle endocytosis | GO:0048488 |  | 4.699E-05 | 0.01615165 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 5.87E-05 | 0.01765296 |
|---|
| regulation of peptidyl-tyrosine phosphorylation | GO:0050730 |  | 6.545E-05 | 0.01749691 |
|---|
| entry into host cell | GO:0030260 |  | 8.596E-05 | 0.02068258 |
|---|
| cGMP metabolic process | GO:0046068 |  | 1.015E-04 | 0.02220473 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| negative regulation of JAK-STAT cascade | GO:0046426 |  | 6.034E-08 | 1.388E-04 |
|---|
| negative regulation of protein kinase cascade | GO:0010741 |  | 4.414E-06 | 5.076E-03 |
|---|
| negative regulation of peptidyl-tyrosine phosphorylation | GO:0050732 |  | 1.508E-05 | 0.01156371 |
|---|
| regulation of JAK-STAT cascade | GO:0046425 |  | 1.924E-05 | 0.01106041 |
|---|
| establishment of tissue polarity | GO:0007164 |  | 2.261E-05 | 0.0103994 |
|---|
| synaptic vesicle endocytosis | GO:0048488 |  | 3.163E-05 | 0.01212338 |
|---|
| regulation of peptidyl-tyrosine phosphorylation | GO:0050730 |  | 6.541E-05 | 0.02149151 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 6.762E-05 | 0.01943946 |
|---|
| platelet-derived growth factor receptor signaling pathway | GO:0048008 |  | 6.762E-05 | 0.01727952 |
|---|
| entry into host cell | GO:0030260 |  | 6.762E-05 | 0.01555157 |
|---|
| ER-associated protein catabolic process | GO:0030433 |  | 8.258E-05 | 0.01726635 |
|---|
| Name | Accession Number | Link | P-val | Corrected P-val |
|---|
| negative regulation of JAK-STAT cascade | GO:0046426 |  | 6.034E-08 | 1.388E-04 |
|---|
| negative regulation of protein kinase cascade | GO:0010741 |  | 4.414E-06 | 5.076E-03 |
|---|
| negative regulation of peptidyl-tyrosine phosphorylation | GO:0050732 |  | 1.508E-05 | 0.01156371 |
|---|
| regulation of JAK-STAT cascade | GO:0046425 |  | 1.924E-05 | 0.01106041 |
|---|
| establishment of tissue polarity | GO:0007164 |  | 2.261E-05 | 0.0103994 |
|---|
| synaptic vesicle endocytosis | GO:0048488 |  | 3.163E-05 | 0.01212338 |
|---|
| regulation of peptidyl-tyrosine phosphorylation | GO:0050730 |  | 6.541E-05 | 0.02149151 |
|---|
| regulation of long-term neuronal synaptic plasticity | GO:0048169 |  | 6.762E-05 | 0.01943946 |
|---|
| platelet-derived growth factor receptor signaling pathway | GO:0048008 |  | 6.762E-05 | 0.01727952 |
|---|
| entry into host cell | GO:0030260 |  | 6.762E-05 | 0.01555157 |
|---|
| ER-associated protein catabolic process | GO:0030433 |  | 8.258E-05 | 0.01726635 |
|---|


